Histone post-translational modifications (PTMs) play crucial roles in chromatin regulation, interacting with a wide array of enzymes, reader proteins, and multi-subunit complexes to coordinate downstream processes. Historically, modified histone peptides have been the favored substrate for in vitro chromatin studies, under the assumption that PTMs are unaffected by higher order nucleosome structure. However, recent work from EpiCypher points to an integral role for nucleosome conformation in dictating histone PTM interactions and chromatin function (Marunde et al. 2022; Morgan et al. 2021).
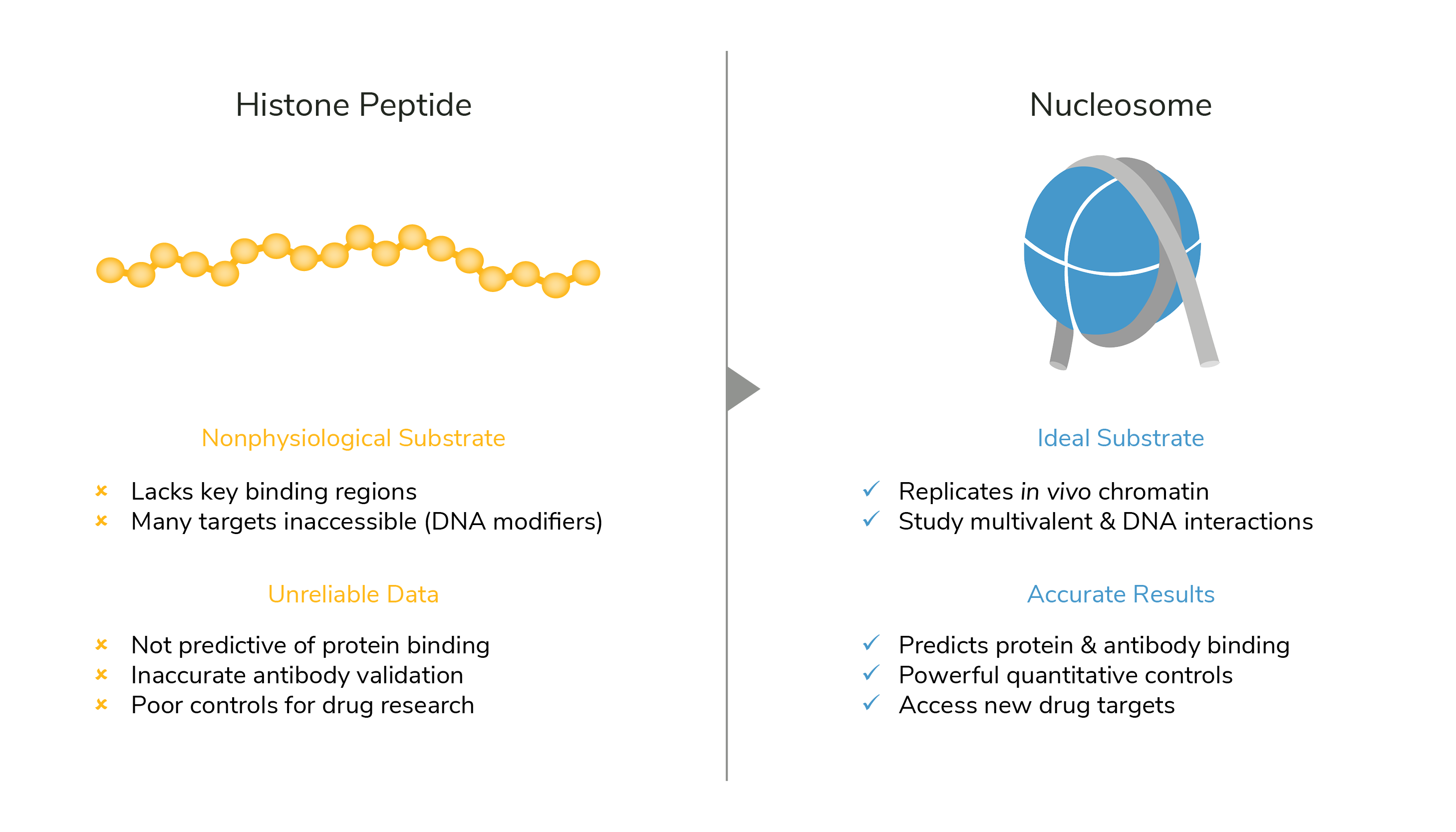
EpiCypher offers a diverse array of recombinant nucleosomes carrying various histone PTMs, histone variants, oncogenic mutations, and more. We also offer custom nucleosome development services and dCypher screening services for specialized research projects and studying chromatin interactions. The use of nucleosome substrates is essential to generate accurate, physiological data, and are highly recommended compared to histone peptides (see Figure below). However, if you would like to still obtain histone peptides, or are interested in the peptides we used to sell, you can contact the UNC Peptide Core Facility that specializes in custom histone peptide synthesis.
Learn more:
- Finding the Best Substrate for Studying Histone Modifications
- Deciphering the Histone Code with the dCypher™ Platform
- The Nucleosome Acidic Patch: A Master Landing Dock for Chromatin Regulators
- Learn More About EpiCypher’s Custom Nucleosome Assembly Services!
- Choosing the Right ChIP Antibody for Your Experiment
- Histone Peptides vs. Nucleosomes : Which is better for ChIP antibody validation?
