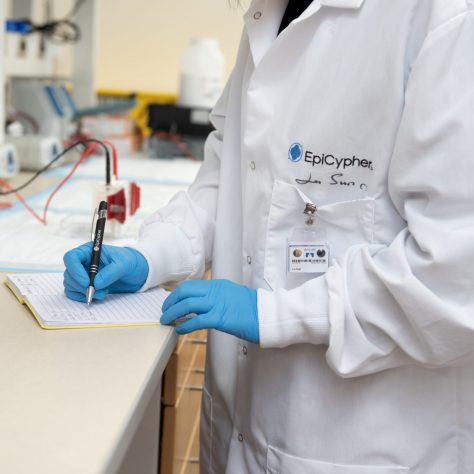
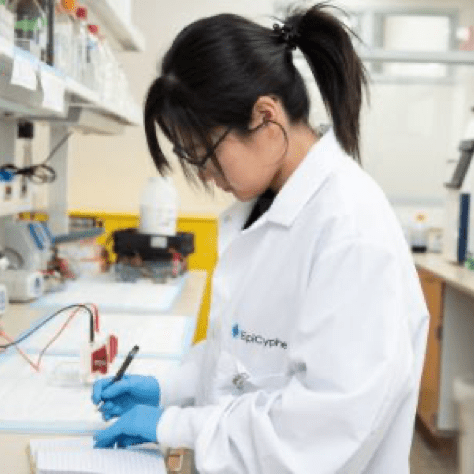
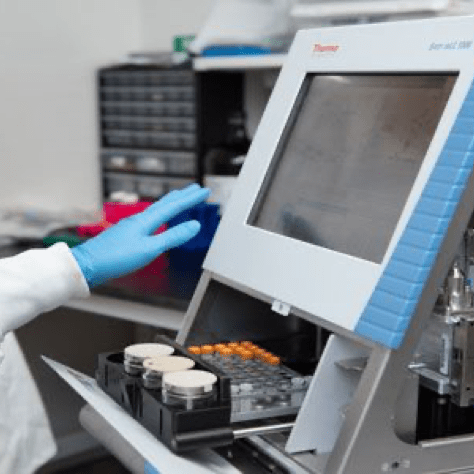
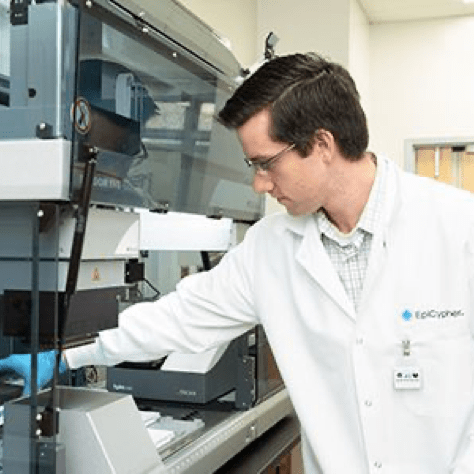
CUTANA™ CUT&RUN Services
Your partner for premium chromatin mapping services
CUT&RUN is the leading technology for mapping genomic enrichment of chromatin targets, such as transcription factors and histone PTMs. EpiCypher’s CUTANA™ CUT&RUN Services provide unique access to our genomic expertise, enabling diverse applications across biomedical research and drug development. As our partner, you can expect:
- Detailed, end-to-end experimental design
- Expert optimization for targets, cell types, tissues, and clinical samples
- Reliable profiling using automated assays and standardized controls
- A comprehensive data report and presentation by our scientists
- Scalable protocols from pilot tests to high-throughput studies
Have Questions?
We’re here to help. Click below and a member of our team will get back to you shortly!
dozens
of partners
Featured Customers



The perfect partnership: your project & our experts
Introduction
Fill out the form below to connect with our team and determine the best solution for your project. EpiCypher CUT&RUN Services have broad application potential, including experiments with large sample numbers and optimization for new chromatin targets or cell types.
Experimental Design
The CUT&RUN Services Team describes our automated workflow, optimized assay controls, and estimated timelines. We work with you to design a comprehensive Statement of Work (SoW), in which we assess project risks and define key details to help ensure robust CUT&RUN results.

Sample delivery and quality control
Our services team has expertise with a variety of cells, tissues, and clinical samples, allowing us to provide end-to-end support for sample prep. Our rigorous quality checks ensure that only the best materials are used in your CUT&RUN experiment.
CUT&RUN experiment and sequencing
Our automated, 96-well plate workflow enables high-throughput chromatin mapping, even at low cell numbers. At each step, defined controls help ensure high reproducibility and flag substandard reactions, allowing you to be confident in robust CUT&RUN results.
Bioinformatic processing and data delivery
Our unique, in-house bioinformatic pipeline was built by EpiCypher scientists for CUT&RUN. After processing, we provide a detailed report tailored to your projects, review key controls to confirm assay success, and provide the files you need to derive biological insights.
Selected Publications
- Agustinus et al. Epigenetic dysregulation from chromosomal transit in micronuclei. Nature 619, 176-183 (2023). PMID: 37286593.
- Baysoy et al. The interweaved signatures of common-gamma-chain cytokines across immunologic lineages. Journal of Experimental Medicine 220 (2023). PMID: 36976164.
- Asberry et al. Reprogramming landscape highlighted by dynamic transcriptomes in therapy-induced neuroendocrine differentiation. Computational and Structural Biotechnology Journal 20, 5873-5885 (2022). PMID: 36382181.
DNA Methylation Sequencing Services
EpiCypher also offers DNA methylation assay services, providing flexible options for genome-wide 5mC mapping or targeted methylation analysis. These methods are powered by CUTANA™ CUT&RUN technology—making them a natural complement to your chromatin profiling projects.

Genome-Wide DNA Methylation Mapping
meCUT&RUN is a novel enrichment-based method for DNA methylation ) sequencing that delivers high-quality, genome-wide profiles at a fraction of the cost and input required for whole-genome bisulfite sequencing.

Targeted DNA Methylation Profiling
Multiomic CUT&RUN is a next-generation epigenomic mapping strategy that profiles chromatin proteins and DNA methylation together in a single reaction, capturing novel insights to drive biological discovery
Tell us about your project:
FAQs about our CUT&RUN Services
Are CUTANA CUT&RUN services right for my project?
CUTANA CUT&RUN chromatin mapping services use automated, quality-controlled workflows to enable medium to large-scale analysis of transcription factors, histone PTMs, and other chromatin-associated targets. Combined with the improved costs and sensitivity of CUT&RUN (compared to ChIP-seq), epigenomic mapping can now be applied at a greater scale than ever before. From early-stage pilot studies requiring optimization for new sample types and targets, to large-scale multi-center collaborations, or drug screens profiling thousands of samples, our genomics experts can help find solutions.
If you are interested, fill out the form above. Our experts can help you estimate total project size and determine if CUT&RUN Services is right for you. If you have a smaller project and/or the capability to bring CUT&RUN in-house, we offer user-friendly CUTANA CUT&RUN and Library Prep Kits, which include all reagents and controls needed for 48 CUT&RUN reactions.
Please note: CUT&RUN Services are for Research Use Only (RUO) as our laboratory is not CLIA certified.
What tissues or cells types are compatible with CUTANA CUT&RUN Services?
Our CUT&RUN Service protocols are optimized for freshly isolated and frozen cells or nuclei from various sources. For certain targets, such as labile histone lysine acetylation PTMs or acetyl-binding proteins, light cross-linking may be recommended. EpiCypher has successfully used our automated CUT&RUN assays for a variety of sample types and cell preparations, including:
- Suspension and adherent cell lines
- Drug-treated or stimulated cells
- Primary cells, including immune cells
- FACS-isolated cells
- Samples generated from tissue (including frozen patient biopsies)
- Clinical samples (PBMCs, BMMCs)
We provide detailed sample prep protocols and shipment instructions. For tissues, we can provide expert guidance on the best strategies for isolating cells.
Unfortunately, our CUT&RUN processes are not yet optimized for fixed tissues embedded in paraffin blocks (e.g. FFPE). Contact us to learn more about ongoing research efforts or alternate offerings to support this application.
Technical How many cells do I need for CUTANA CUT&RUN Services?
For CUTANA CUT&RUN experiments, we recommend using 500,000 to 50,000 cells per reaction.*
Note that each reaction maps one target in one sample. Samples are unique cell populations, such as control vs. mutant or biological replicates.
If you have one cell sample, and you want to map three unique chromatin targets, you will be setting up three reactions – and thus require 1.5 million cells. We always suggest collecting 10-20% excess cells to account for sample loss.
* Our workflow is validated down to 10,000 cells for select targets. Success at low cell numbers depends on antibody performance, target abundance, and sample quality. There are additional risks assumed by going below the recommended cell numbers – such as low yields, increased adapter dimer contamination in libraries, increased background and/or sequencing depth – that may require additional optimization. Our scientists will help you weigh these risks during the experimental design phase of the project.
What controls are included?
Controls are essential for overall project success, risk mitigation, and data interpretation. For this reason, EpiCypher’s CUT&RUN Services workflow includes a collection of quality control checks at each step of the process to help ensure you get the highest quality data possible.
Note that in our workflow, samples are unique cell populations, such as control vs. mutant or biological replicates. The term reaction is used to define the mapping of one target in one unique sample. For instance, if you have one cell sample and want to map three distinct chromatin targets, you will have a total of 3 CUT&RUN reactions.
Sample preparation
- Detailed methods for cell or nuclei collection and shipping instructions are provided to promote seamless sample handoff to our team.
- For tissues, our team can help develop a standard procedure for cell harvest.
- Upon arrival at EpiCypher, our team confirms cell number, viability, and integrity. Our scientists use these data to create a sample report that we review together to assess risk.
- Only samples that pass quality control metrics are included in experiments.
Control reactions including SNAP-CUTANA™ Spike-in Controls
- We set up three control reactions for each unique sample: H3K4me3 and H3K27me3 (positive controls) and IgG (negative controls). Example: If your project analyzes 5 unique PBMC samples, we set up three control reactions for each sample, or a total of 15 control reactions.
- Control reactions are used to validate sample quality, examine assay background, determine significant regions of enrichment above background (e.g. peaks), and confirm workflow success.
- Additionally, data from positive controls provide valuable biological context for chromatin regions associated with active (H3K4me3) and repressed (H3K27me3) genes.
- As an additional layer of protection, control reactions include our SNAP-CUTANA Spike-in Controls. SNAP-CUTANA Spike-ins comprise a defined, proprietary control that is spiked into reactions at the beginning of the experiment. The control is processed alongside biological samples through every step of the protocol, enabling direct monitoring and readout of experimental success. Learn more about our SNAP-CUTANA Spike-in Technology here.
E. coli Spike-in DNA
- All reactions are spiked with E. coli Spike-in DNA after targeted MNase cleavage to monitor library prep success.
- E. coli read counts are provided to customers, which can be used for sequencing normalization.
Pre-sequencing metrics
- EpiCypher scientists examine raw CUT&RUN-enriched DNA yields.
- Adjustments are made for low yields and/or low-abundant targets when going into library prep.
- Fragment distribution and concentration of sequencing libraries are determined using the Agilent TapeStation system. This is the best method to confirm CUT&RUN success before sequencing.
Post-sequencing metrics
- Libraries are sequenced to a depth that provides >3 million uniquely mapped reads. EpiCypher scientists use an optimized bioinformatic pipeline to align sequencing reads to the respective species genome and deliver high-quality data.
- We examine overall sequencing statistics, including unique alignment rates, duplication rates, and percent of reads assigned to E. coli and SNAP-CUTANA Spike-in controls, all of which help to determine sequencing success.
- Genomic and SNAP-CUTANA spike-in sequences are examined for control reactions (IgG, H3K4me3, H3K27me3) to ensure they produce the expected biological distribution with high specificity and signal-to-noise. This analysis lends confidence that the samples were high quality and the overall experiment was successful, enabling confident interpretation of the target(s) of interest.
- Peak calling pipelines are used to identify areas of significant target enrichment and assess overall signal-to-noise (Fraction of Reads in Peaks, or FRiP score).
- If multiple CUT&RUN experimental variables (e.g. antibodies, sample prep methods, etc.) are tested, the metrics above will define the optimal conditions for downstream analyses and future projects.
What targets can I map? Can you help me find a good antibody?
The EpiCypher Services Team has experience mapping various targets:
- Transcription factors (e.g. CTCF, FOXA1, p53, c-Jun)
- Chromatin remodeling enzymes (e.g. SMARCA2/BRM and SMARCA4/BRG1)
- Histone modifying enzymes and regulators (e.g. MLL1, EZH2, BRD4, Menin)
- Histone PTMs (methylation, acetylation, phosphorylation, ubiquitination)
As a result, we have a broad catalog of target-specific antibodies demonstrating exceptional performance in CUT&RUN. We are also highly skilled in sourcing and validating antibodies to new targets. Antibody screening is offered as part of our CUT&RUN Services and can be incorporated into your Statement of Work.
What data do I receive?
We work with every service partner to tailor our analysis to your needs. EpiCypher uses our bioinformatics pipeline – specifically optimized for CUT&RUN – to analyze sequencing data. At our data delivery meeting, we review preliminary data visualization at customer-identified target loci and all quality control checks (outlined above) to illustrate experimental success. We coordinate handoff of raw sequencing data, alignment files, bigWig files, and called peaks by SharePoint or Amazon Web Services. Files can be immediately used to visualize data, perform downstream analyses (e.g. differential enrichment), and derive biological insights in your lab. Specifically, we provide:
- Raw sequencing data in FASTQ files. EpiCypher performs paired-end sequencing, meaning each sequencing library has two FASTQ files (e.g. R1 and R2).
- Aligned, filtered reads in BAM files. The final BAM files contain uniquely aligned reads, with multimapping reads, exclusion list regions, and duplicate reads removed.
- Visualization-ready data in bigWig files. BigWig files are RPKM (Reads per Kilobase per Million mapped reads) normalized for visualization on genomic browsers (e.g. IGV).
- Signal enrichment in BED files, generated from peak calling.
- (As needed) Text file with motif analysis results, showing target-enriched sequences.
To see a sample data report, email [email protected]




