Choosing the Right ChIP Antibody for Your Experiment
- Ellen Weinzapfel

There are multiple components to designing and executing a successful chromatin immunoprecipitation (ChIP) or ChIP-seq experiment:
- – What target should I study (i.e. histone post-translational modification [PTM] or a chromatin-associated protein, such as a transcription factor)?
- – What cell line or tissue type should I use?
- – What ChIP method is best (e.g. cross-linked or native)?
However, the single most important step you can take in ensuring your experiment will yield high-quality data is choosing the right ChIP antibody – as the most well-optimized protocol will still fail if you are using a low performance antibody! Here are some major points to consider when shopping for an antibody to run your next ChIP experiment against histone PTMs.
ChIP Antibody Specificity
It is crucial to make sure that your antibody specifically recognizes its designated target. This is true for ALL ChIP experiments, but is particularly important in the study of histone PTMs, as we (and others) have shown that antibodies to PTMs exhibit vastly different binding capabilities 1-5.
Modified histone peptide arrays have long been the gold standard in validating ChIP antibody specificity to histone PTMs. However, they fail to accurately model endogenous chromatin structures and do not recapitulate binding conditions in ChIP assays. As we described in our previous blog post, histone peptide arrays are a poor predictor of binding activity in ChIP, and have had massive and negative impacts on the quality of histone PTM antibodies 5.
The best way to determine antibody specificity, for any target, is to test its binding activity against physiologically relevant controls in the context of your experiment.
To address these problems and provide a path forward for superior ChIP studies, EpiCypher SNAP-ChIP® Certified histone PTM antibodies are validated against a panel of on- and off-target recombinant nucleosomes, which are added to bulk chromatin prior to immunoprecipitation. These SNAP-ChIP® spike-ins enable in situ monitoring of antibody specificity, binding efficiency (see more on this below), and technical variation, providing unprecedented user control over ChIP experiments.
Why does this matter? As we detailed in our previous blog post, SNAP-ChIP has revealed vast differences in specificity for antibodies targeting the same histone PTM (see Figure 1). In our recent study of H3K4me1, me2, and me3 antibodies, we found that the most highly cited H3K4me3 ChIP antibody displayed >50% cross reactivity with H3K4me2 (Figure 1) 5. These findings have been corroborated by an independent study 6, and suggest that poor antibody validation may lead to incorrect biological interpretations.
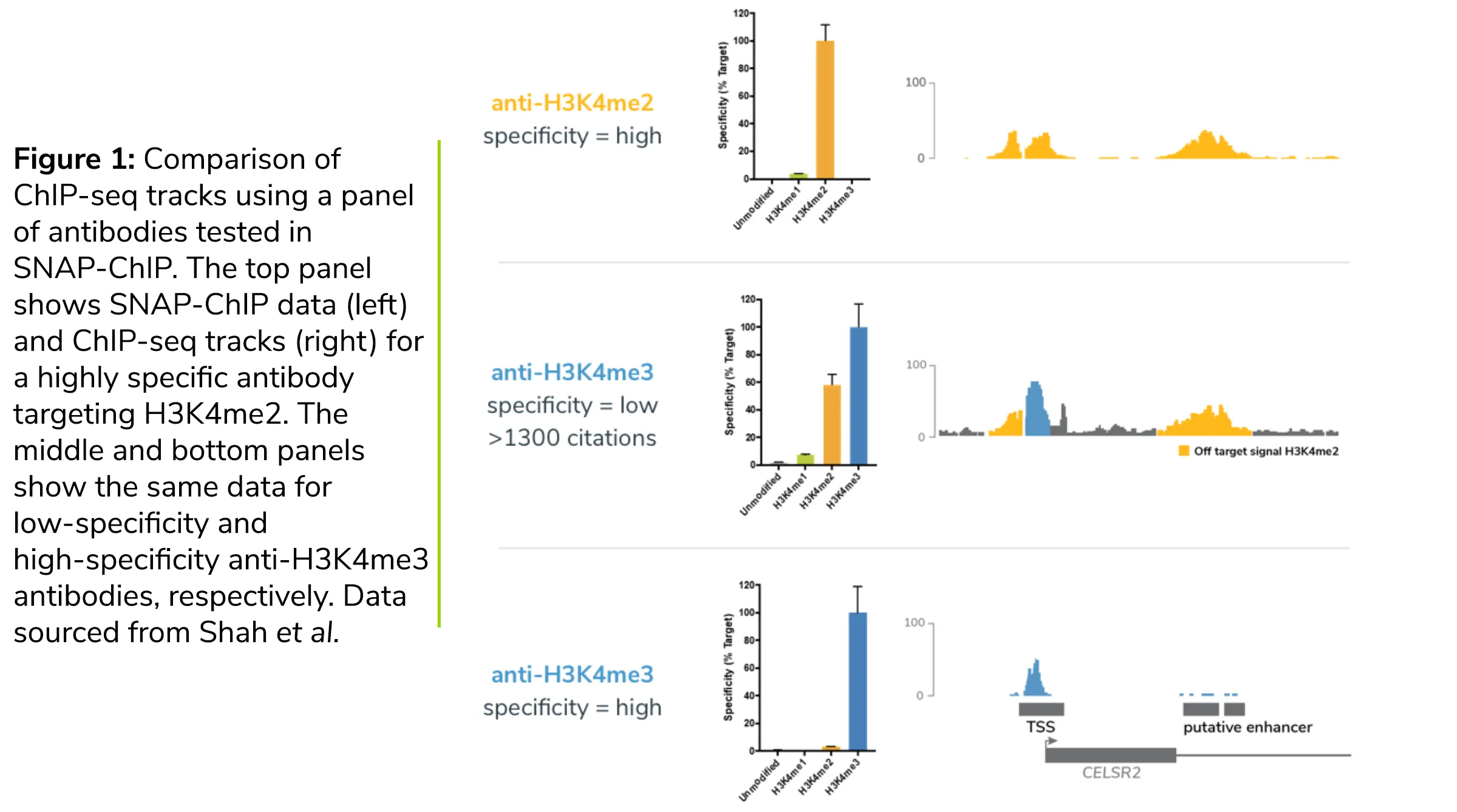
ChIP Antibody Enrichment (IP Efficiency)
The efficiency of an antibody in ChIP refers to the enrichment of the target following IP. Typically, enrichment is determined by performing qPCR for a positive control locus, calculating the signal from the ChIP reaction as a percent of the signal from Input chromatin. However, if the relative abundance of the epitope in bulk chromatin is unknown, this information may not be very useful, or accurate.
SNAP-ChIP validation provides a quantitative metric for efficiency, by calculating the recovery of on-target SNAP-ChIP nucleosomes that were spiked into bulk chromatin. To pass SNAP-ChIP certification testing, the antibody must recover at least 5% of the spike-in nucleosome.
We have found that a high antibody efficiency correlates with increased sample recovery (i.e. more material to product high quality sequencing libraries) and improved signal-to-noise (S/N) in ChIP (see Figure 2).

Clonal Variation
The last thing we will consider are potential lot-to-lot changes in ChIP antibody performance. Scientists commonly validate an antibody the first time they purchase it, and if it passes this testing they will often continue to reorder when it runs out, without worrying about recertification.
However, these practices assume that antibodies remain relatively the same in binding affinity and specificity over time.
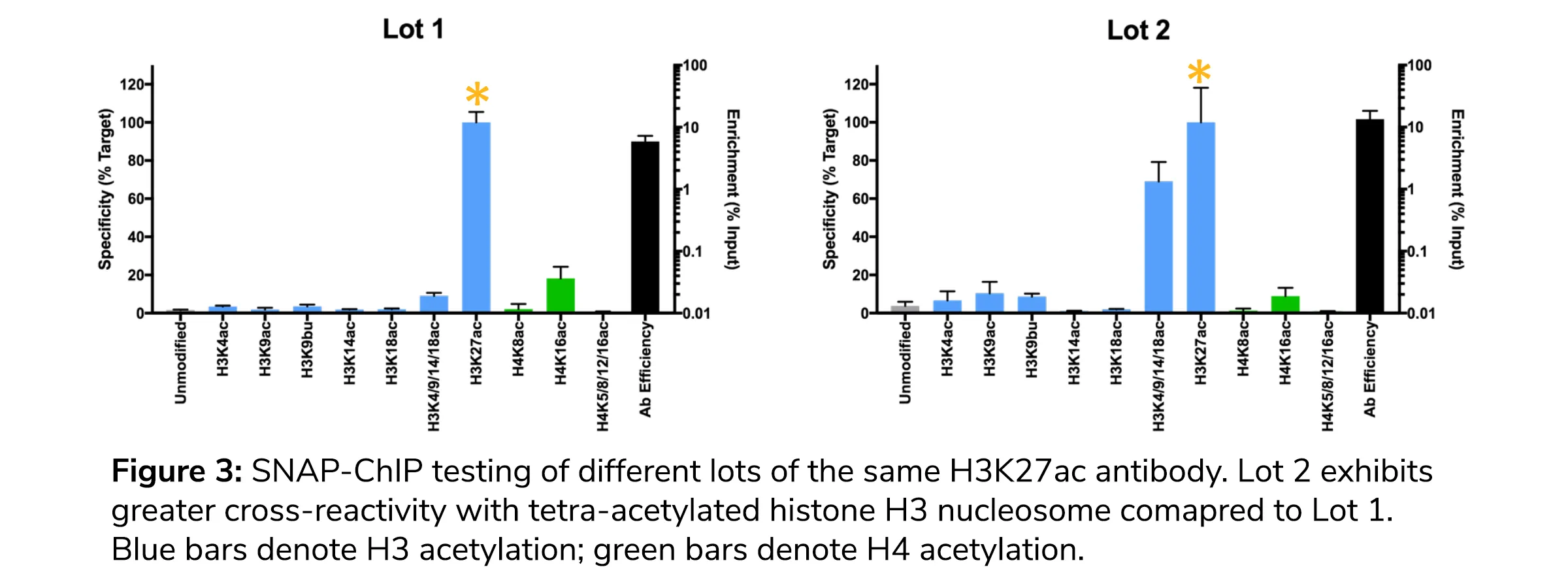
Why does this matter? Our internal research suggests that there can be substantial changes in antibody performance between production lots, both in terms of cross-reactivity and on-target PTM enrichment (see Figure 3). Thus, scientists must remain vigilant with their antibodies and revalidate their ChIP antibody every time they purchase a new lot.
Summary and future multiomic CUT&Tag applications
CUT&Tag, with its ability to profile multiple histone modifications and other chromatin features alongside transcriptome profiling, has become a key tool in multiomics. Its adaptability when combined with single-cell technologies offers unprecedented insights into the epigenetic landscape and the dynamic interplay between chromatin states and gene expression.
Future integrations of CUT&Tag will provide deeper insights into regulatory mechanisms governing cellular functions and disease development. For example, the development of CUT&Tag-based chromatin accessibility assays (CUTAC) has allowed researchers to probe histone modifications and active regulatory regions in FFPE-embedded tissues18,19, a feat that has been incredibly difficult to accomplish even with highly sensitive ATAC-seq approaches. CUT&Tag has also been leveraged in several studies for spatial profiling, which preserves tissue architecture in addition to providing high-resolution, single-cell chromatin profiling – see this blog to learn more.
Of course, multiomics encompasses dozens of other strategies, and many have been paired with ATAC-seq or Tn5 in previous papers. Drawing on these reports, other CUT&Tag multiomic applications include:
- DNA methylation (e.g. ATAC-Me19)
- Spatial profiling (e.g. Spatial CUT&Tag20)
- Protein levels (e.g. PHAGE-ATAC21)
- Lineage tracing22
A New Way Forward : SNAP-ChIP® Certified Antibodies
EpiCypher’s SNAP-ChIP technology provides the first method to determine antibody specificity and efficiency in your ChIP experiment. These in situ controls provide a powerful new platform to examine assay performance before you spend your precious funds on sequencing.
To deliver the best quality antibodies on the market, EpiCypher has begun leveraging SNAP-ChIP technology to identify truly fit-for-purpose antibodies. To date, we have screened >400 antibodies using SNAP-ChIP to identify best-in-class antibodies to a wide collection of histone PTMs, including:
<ul “=””>
- – lysine methylation states (mono-, di-, and tri-methylation),
- – lysine acylation (acetylation as well as extended acyl states, including butyrylation and crotonylation),
- – arginine methylation (mono-, di-symmetric, and di-asymmetric),
- – arginine citrullination
- – oncogenic histone mutations.
We are now offering a widely diverse list of antibodies that pass our rigorous SNAP-ChIP Certification criteria (high specificity & efficiency). For the most updated list of SNAP-ChIP certified antibodies, click here.
Our partners at ThermoFisher Scientific also have an extensive collection of ChIP-grade antibodies against multiple targets, and have implemented SNAP-ChIP testing for many of their histone PTM antibodies! Click here for more info. If your PTM of interest is not on either list, please inquire at [email protected] for personalized support from our experienced scientists!
Questions or comments? Let us know!
References
1. Rothbart SB, et al. Peptide microarrays to interrogate the “histone code”. Methods Enzymol, 2012. 512:p. 107-35. (PubMed PMID: 22910205) (PMC3741997)
2. Rothbart SB, et al. Poly-acetylated chromatin signatures are preferred epitopes for site-specific histone H4 acetyl antibodies. Sci Rep, 2012. 2: p. 489. (PubMed PMID: 22761995) (PMC3388470)
3. Rothbart SB, et al. An Interactive Database for the Assessment of Histone Antibody Specificity. Mol Cell, 2015. 59(3): p. 502-11.(PubMed PMID: 26212453) (PMC4530063)
4. Cornett EM, et al. Analysis of Histone Antibody Specificity with Peptide Microarrays. J Vis Exp, 2017. (126): p. (PubMed PMID: 28809825)
5. Shah RN, et al. Examining the Roles of H3K4 Methylation States with Systematically Characterized Antibodies. Mol Cell, 2018. 72(1): p. 162-77 e7. ( PubMed PMID: 30244833(PMC6173622)
6. Lam KG, et al. Cell-type-specific genomics reveals histone modification dynamics in mammalian meiosis. Nat Commun, 2019. 10(1): p. 3821. ( PubMed PMID: 31444359 ) (PMC6707301)
