ELF1 CUTANA™ CUT&RUN Antibody
{"url":"https://www.epicypher.com/products/antibodies/cutana-cut-run-antibodies/cut-run-antibodies-chromatin-associated-proteins/elf1-cutana-cut-run-antibody","add_this":[{"service":"facebook","annotation":""},{"service":"email","annotation":""},{"service":"print","annotation":""},{"service":"twitter","annotation":""},{"service":"linkedin","annotation":""}],"gtin":null,"id":907,"bulk_discount_rates":[],"can_purchase":true,"meta_description":"Rabbit polyclonal ELF1 antibody rigorously tested for robust and reliable performance in CUT&RUN assays. Also validated for IHC.","category":["Chromatin Mapping/CUTANA™ ChIC / CUT&RUN Assays","Antibodies/CUTANA™ CUT&RUN Antibodies","Antibodies/CUTANA™ CUT&RUN Antibodies/CUT&RUN Antibodies - Chromatin-Associated Proteins"],"AddThisServiceButtonMeta":"","main_image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/907/1009/chd3-cutana-cutandrun-antibody__66975.1645734525__91145.1648666822__83973.1648670498.jpg?c=2","alt":"ELF1 CUTANA™ CUT&RUN Antibody"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=907","shipping":{"calculated":true},"num_reviews":0,"weight":"0.01 LBS","custom_fields":[{"id":"1029","name":"Pack Size","value":"100 µL"}],"sku":"13-2023","description":"<div class=\"product-general-info\">\n <ul class=\"product-general-info__list-left\">\n <li class=\"product-general-info__list-item\">\n <strong>Type: </strong>Polyclonal\n </li>\n <li class=\"product-general-info__list-item\">\n <strong>Host: </strong>Rabbit\n </li>\n <li class=\"product-general-info__list-item\">\n <strong>Applications: </strong>CUT&RUN, IHC\n </li>\n </ul>\n <ul class=\"product-general-info__list-right\">\n <li class=\"product-general-info__list-item\">\n <strong>Reactivity: </strong>Human\n </li>\n <li class=\"product-general-info__list-item\">\n <strong>Format: </strong>Antigen affinity-purified\n </li>\n <li class=\"product-general-info__list-item\">\n <strong>Target Size: </strong>67 kDa\n </li>\n </ul>\n</div>\n\n<div class=\"service_accordion product-droppdown\">\n <div class=\"container\">\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel current\">\n <h3 class=\"sub-title1\">Description</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description-specific\">\n <p>\n This antibody meets EpiCypher’s \"CUTANA Compatible\" criteria for\n performance in Cleavage Under Targets and Release Using Nuclease\n (CUT&RUN) and/or Cleavage Under Targets and Tagmentation (CUT&Tag)\n approaches to genomic mapping. Every lot of a CUTANA Compatible\n antibody is tested in the indicated approach using EpiCypher\n optimized <a href=\"/resources/protocols/\">protocols</a> and\n determined to yield peaks that show a genomic distribution pattern\n consistent with reported function(s) of the target protein. ELF1 is\n a member of the ETS (e26 transformation specific sequence) family of\n transcription factor proteins. ELF1 antibody produces CUT&RUN peaks\n above background primarily in intronic and promoter regions (<strong\n >Figure 1</strong\n >) that overlap with known ELF1 DNA-binding motifs (<strong\n >Figure 2</strong\n >). ELF1 is expressed in B and T cells and has been shown to be\n involved with the regulation of several T- and B-cell specific genes\n [1].\n </p>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel current\">\n <h3 class=\"sub-title1\">Validation Data</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description-specific\">\n <p>\n CUT&RUN was performed on 500k HeLa cells with 0.5 µg of either ELF1\n or IgG negative control (EpiCypher\n <a\n href=\"/products/nucleosomes/snap-cutana-spike-in-controls/cutana-rabbit-igg-cut-run-negative-control-antibody\"\n >13-0042</a\n >) antibodies using the CUTANA™ ChIC/CUT&RUN Kit v2.0 (EpiCypher\n <a\n href=\"/products/epigenetics-reagents-and-assays/cutana-chic-cut-and-run-kit\"\n >14-1048</a\n >). Library preparation was performed with 5 ng of DNA (or the total\n amount recovered if less than 5 ng) using the CUTANA™ CUT&RUN\n Library Prep Kit (EpiCypher\n <a\n href=\"/products/epigenetics-reagents-and-assays/cutana-cut-and-run-library-prep-kit\"\n >14-1001/14-1002</a\n >). Both kit protocols were adapted for high throughput Tecan liquid\n handling. Libraries were run on an Illumina NextSeq2000 with\n paired-end sequencing (2x50 bp). Sample sequencing depth was 5.0\n million reads (IgG) and 6.0 million reads (ELF1). Data were aligned\n to the hg19 genome using Bowtie2. Data were filtered to remove\n duplicates, multi-aligned reads, and ENCODE DAC Exclusion List regions.\n </p>\n\n <section class=\"image-picker\">\n <div class=\"image-picker__left\">\n <div\n class=\"image-picker__main-content_active image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg\n class=\"image-picker__svg-left\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\">\n <path\n d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a\n href=\"/content/images/products/antibodies/13-2023-elf1-peaks.jpeg\"\n target=\"_blank\"\n class=\"image-picker__main-image-link\"\n ><img loading=\"lazy\"\n alt=\"13-2023-elf1-peaks\"\n src=\"/content/images/products/antibodies/13-2023-elf1-peaks.jpeg\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\"\n >(Click to enlarge)</span\n ></a\n >\n <button class=\"image-picker__right-arrow\">\n <svg\n class=\"image-picker__svg-right\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\">\n <path\n d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\"\n ><strong>Figure 1: ELF1 peaks in CUT&RUN. </strong><br />\n CUT&RUN was performed as described above. Peaks were called\n using MACS2. Heatmaps show ELF1 peaks relative to IgG\n negative control antibody in aligned rows ranked by\n intensity (top to bottom) and color such that red indicates\n high localized enrichment and blue denotes background signal\n (<strong>left</strong>). The number of peaks that fall into\n distinct classes of functionally annotated genomic regions\n are shown (<strong>right</strong>).\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg\n class=\"image-picker__svg-left\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\">\n <path\n d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a\n href=\"/content/images/products/antibodies/13-2023-binding-motif-analysis.jpeg\"\n target=\"_blank\"\n class=\"image-picker__main-image-link\"\n ><img loading=\"lazy\"\n alt=\"13-2023-binding-motif-analysis\"\n src=\"/content/images/products/antibodies/13-2023-binding-motif-analysis.jpeg\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\"\n >(Click to enlarge)</span\n ></a\n >\n <button class=\"image-picker__right-arrow\">\n <svg\n class=\"image-picker__svg-right\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\">\n <path\n d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\"\n ><strong\n >Figure 2: ELF1 transcription factor binding motif\n analysis in CUT&RUN </strong\n ><br />\n Homer analysis determined that the ELF1(ETS)/Jurkat-ELF1-\n ChIP-Seq(SRA014231)/Homer consensus motif, represented as a\n sequence logo position weight matrix, was significantly\n enriched under ELF1 CUT&RUN peaks (<strong>left</strong>).\n The number of ELF1 peaks containing the consensus motif from\n the left panel is represented by a Venn Diagram\n (<strong>right</strong>).\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg\n class=\"image-picker__svg-left\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\">\n <path\n d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a\n href=\"/content/images/products/antibodies/13-2023-representative-browser-tracks.jpeg\"\n target=\"_blank\"\n class=\"image-picker__main-image-link\"\n ><img loading=\"lazy\"\n alt=\"13-2023-representative-browser-tracks\"\n src=\"/content/images/products/antibodies/13-2023-representative-browser-tracks.jpeg\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\"\n >(Click to enlarge)</span\n ></a\n >\n <button class=\"image-picker__right-arrow\">\n <svg\n class=\"image-picker__svg-right\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\">\n <path\n d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\"\n ><strong\n >Figure 3: ELF1 CUT&RUN representative browser tracks </strong\n ><br />\n CUT&RUN was performed as described above. Gene browser shots\n were generated using the Integrative Genomics Viewer (IGV,\n Broad Institute). Three representative loci of the top\n called peaks are shown.\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg\n class=\"image-picker__svg-left\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\">\n <path\n d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a\n href=\"/content/images/products/antibodies/13-2023-immunohistochemistry-data.jpeg\"\n target=\"_blank\"\n class=\"image-picker__main-image-link\"\n ><img loading=\"lazy\"\n alt=\"13-2023-immunohistochemistry-data\"\n src=\"/content/images/products/antibodies/13-2023-immunohistochemistry-data.jpeg\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\"\n >(Click to enlarge)</span\n ></a\n >\n <button class=\"image-picker__right-arrow\">\n <svg\n class=\"image-picker__svg-right\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\">\n <path\n d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\"\n ><strong>Figure 4: Immunohistochemistry data </strong><br />\n FFPE sections of human Hodgkin’s Lymphoma\n (<strong>left</strong>) and human ovarian carcinoma\n (<strong>right</strong>) using ELF1 antibody at a dilution\n of 1:250.\n </span>\n </p>\n </div>\n </div>\n <aside class=\"image-picker__right\">\n <div class=\"image-picker__gallery\">\n <img loading=\"lazy\"\n alt=\"13-2023-elf1-peaks\"\n src=\"/content/images/products/antibodies/13-2023-elf1-peaks.jpeg\"\n width=\"200\"\n class=\"image-picker__side-image image-picker__side-image_active\"\n role=\"button\" />\n <img loading=\"lazy\"\n alt=\"13-2023-binding-motif-analysis\"\n src=\"/content/images/products/antibodies/13-2023-binding-motif-analysis.jpeg\"\n width=\"200\"\n class=\"image-picker__side-image image-picker__side-image_active\"\n role=\"button\" />\n <img loading=\"lazy\"\n alt=\"13-2023-representative-browser-tracks\"\n src=\"/content/images/products/antibodies/13-2023-representative-browser-tracks.jpeg\"\n width=\"200\"\n class=\"image-picker__side-image image-picker__side-image_active\"\n role=\"button\" />\n <img loading=\"lazy\"\n alt=\"13-2023-immunohistochemistry-data\"\n src=\"/content/images/products/antibodies/13-2023-immunohistochemistry-data.jpeg\"\n width=\"200\"\n class=\"image-picker__side-image image-picker__side-image_active\"\n role=\"button\" />\n </div>\n </aside>\n </section>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Technical Information</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\">\n <div class=\"product-tech-info\">\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Immunogen</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n Between amino acids 569 and 619\n </div>\n </div>\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Storage</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n Stable for 1 year at 4°C from date of receipt\n </div>\n </div>\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Formulation</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n Antigen affinity-purified antibody in Tris-buffered saline, 0.1%\n BSA, 0.9% sodium azide\n </div>\n </div>\n </div>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Recommended Dilution</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\">\n <div class=\"product-tech-info\">\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>CUT&RUN:</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n 0.5 µg per reaction\n </div>\n </div>\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Immunohistochemistry (IHC):</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n 1:100 - 1:500\n </div>\n </div>\n </div>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Gene & Protein Information</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\">\n <div class=\"product-tech-info\">\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>UniProt ID</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">P32519</div>\n </div>\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Gene Name</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n E74 like ETS transcription factor 1\n </div>\n </div>\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Protein Name</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n ETS-related transcription factor Elf-1\n </div>\n </div>\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Alternate Names</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n ETS-like factor 1, EFTUD1, RIA1\n </div>\n </div>\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Target Size</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">67 kDa</div>\n </div>\n </div>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">References</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\">\n <strong>Background References:</strong>\n <br />\n [1] Oettgen et al. <em>Mol Cell Biol</em> (1996). PMID:\n <a\n href=\"https://pubmed.ncbi.nlm.nih.gov/8756667/\"\n title=\"Characterization of NERF, a novel transcription factor related to the Ets factor ELF-1\"\n target=\"new\">\n 8756667</a\n ><br />\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Documents & Resources</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\">\n <div class=\"product-documents\">\n <a\n href=\"/content/documents/tds/13-2023.pdf\"\n target=\"_blank\"\n class=\"product-documents__link\">\n <svg\n version=\"1.1\"\n id=\"Layer_1\"\n xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\"\n x=\"0px\"\n y=\"0px\"\n viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\"\n xml:space=\"preserve\"\n class=\"product-documents__icon\"\n alt=\"16-0030 Datasheet\">\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M191.92,68.77l-47.69-47.69c-1.33-1.33-3.12-2.08-5.01-2.08H45.09C41.17,19,38,22.17,38,26.09v184.36\n c0,3.92,3.17,7.09,7.09,7.09h141.82c3.92,0,7.09-3.17,7.09-7.09V73.8C194,71.92,193.25,70.1,191.92,68.77z M177.65,77.06h-41.7\n v-41.7L177.65,77.06z M178.05,201.59H53.95V34.95h66.92v47.86c0,5.14,4.17,9.31,9.31,9.31h47.86V201.59z\" />\n </g>\n <rect\n x=\"20\"\n y=\"112\"\n class=\"product-documents__svg-background\"\n width=\"146\"\n height=\"76\" />\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M23.83,125.68h22.36c5.29,0,9.41,1.33,12.35,4c2.94,2.67,4.42,6.39,4.42,11.18c0,4.78-1.47,8.51-4.42,11.18\n c-2.94,2.67-7.06,4-12.35,4H34.59v18.29H23.83V125.68z M44.81,147.9c5.38,0,8.07-2.32,8.07-6.97c0-2.39-0.67-4.16-2-5.31\n c-1.33-1.15-3.36-1.73-6.07-1.73H34.59v14.01H44.81z\" />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M69.92,125.68h18.91c5.29,0,9.84,0.97,13.66,2.9c3.82,1.93,6.74,4.72,8.76,8.35\n c2.02,3.63,3.04,7.98,3.04,13.04c0,5.06-1,9.42-3,13.08c-2,3.66-4.91,6.45-8.73,8.38c-3.82,1.93-8.4,2.9-13.73,2.9H69.92V125.68z\n M88.07,165.63c10.35,0,15.52-5.22,15.52-15.66c0-10.4-5.17-15.59-15.52-15.59h-7.38v31.26H88.07z\" />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M122.57,125.68h32.84v8.49h-22.22v11.18h20.84v8.49h-20.84v20.49h-10.63V125.68z\" />\n </g>\n </svg>\n <span class=\"product-documents__info\">Technical Datasheet</span>\n </a>\n </div>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Additional Info</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\">\n <p>\n This antibody is provided for commercial sale under license from\n Bethyl Laboratories, Inc.\n </p>\n </div>\n </div>\n </div>\n </div>\n</div>\n","tags":[],"warranty":"","price":{"without_tax":{"formatted":"$525.00","value":525,"currency":"USD"},"tax_label":"Sales Tax"},"detail_messages":"","availability":"","page_title":"ELF1 CUTANA™ CUT&RUN Antibody","cart_url":"https://www.epicypher.com/cart.php","max_purchase_quantity":0,"mpn":null,"upc":null,"options":[],"related_products":[{"id":889,"sku":null,"name":"CUTANA™ CUT&RUN Library Prep Kit","url":"https://www.epicypher.com/products/epigenetics-kits-and-reagents/cutana-cut-run-library-prep-kit","availability":"","rating":null,"brand":{"name":null},"category":["Chromatin Mapping","Chromatin Mapping/CUTANA™ ChIC / CUT&RUN Assays"],"summary":"\n \n \n \n \n \n \n \n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/889/997/Epicypher_CUTANA_CUTRUN_LibraryPrep_Box__64431.1646406521.png?c=2","alt":"CUTANA™ CUT&RUN Library Prep Kit"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/889/997/Epicypher_CUTANA_CUTRUN_LibraryPrep_Box__64431.1646406521.png?c=2","alt":"CUTANA™ CUT&RUN Library Prep Kit"}],"date_added":"31st Jan 2022","pre_order":false,"show_cart_action":true,"has_options":true,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":983,"name":"Pack Size","value":"48 Reactions"}],"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"price":{"without_tax":{"currency":"USD","formatted":"$1,525.00","value":1525},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=889"},{"id":694,"sku":null,"name":"CUTANA™ pAG-MNase for ChIC/CUT&RUN Workflows","url":"https://www.epicypher.com/products/epigenetics-reagents-and-assays/cutana-pag-mnase-for-chic-cut-and-run-workflows","availability":"","rating":null,"brand":{"name":null},"category":["Chromatin Mapping","Chromatin Mapping/CUTANA™ ChIC / CUT&RUN Assays"],"summary":"\n \n \n Type: Nuclease\n \n \n Mol Wgt: 43.7 kDa\n \n \n \n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/694/689/Screen_Shot_2020-02-12_at_11.01.55_AM__17144.1581530752.png?c=2","alt":"CUTANA™ pAG-MNase for ChIC/CUT&RUN Workflows"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/694/689/Screen_Shot_2020-02-12_at_11.01.55_AM__17144.1581530752.png?c=2","alt":"CUTANA™ pAG-MNase for ChIC/CUT&RUN Workflows"}],"date_added":"12th Aug 2019","pre_order":false,"show_cart_action":true,"has_options":true,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":1174,"name":"Internal Comment","value":"Excess in bottom of Venom"},{"id":1175,"name":"Internal Comment","value":"Bulk in Psylocke"}],"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"price":{"without_tax":{"currency":"USD","formatted":"$335.00","value":335},"price_range":{"min":{"without_tax":{"currency":"USD","formatted":"$335.00","value":335},"tax_label":"Sales Tax"},"max":{"without_tax":{"currency":"USD","formatted":"$1,295.00","value":1295},"tax_label":"Sales Tax"}},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=694"},{"id":758,"sku":"13-0042","name":"CUTANA™ IgG Negative Control Antibody for CUT&RUN and CUT&Tag","url":"https://www.epicypher.com/products/nucleosomes/snap-cutana-spike-in-controls/cutana-igg-negative-control-antibody-for-cut-run-and-cut-tag","availability":"","rating":null,"brand":{"name":null},"category":["Nucleosomes/SNAP-CUTANA™ Spike-in Controls","Antibodies/CUTANA™ CUT&RUN Antibodies","Chromatin Mapping","Chromatin Mapping/CUTANA™ ChIC / CUT&RUN Assays","Chromatin Mapping/CUTANA™ CUT&Tag Assays"],"summary":"\n \n \n Type: Polyclonal\n \n \n Host: Rabbit\n \n \n Applications: CUT&R","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/758/739/1579731877.1280.1280__61771.1592491196.png?c=2","alt":"CUTANA™ IgG Negative Control Antibody for CUT&RUN and CUT&Tag"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/758/739/1579731877.1280.1280__61771.1592491196.png?c=2","alt":"CUTANA™ IgG Negative Control Antibody for CUT&RUN and CUT&Tag"}],"date_added":"17th Jun 2020","pre_order":false,"show_cart_action":true,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":1148,"name":"Pack Size","value":"100 µg"},{"id":1149,"name":"Internal Comment","value":"bulk stocks from Thermo in cold room"}],"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"add_to_cart_url":"https://www.epicypher.com/cart.php?action=add&product_id=758","price":{"without_tax":{"currency":"USD","formatted":"$105.00","value":105},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=758"}],"shipping_messages":[],"rating":0,"meta_keywords":"ELF1 antibody, CUT&RUN antibody, transcription factor antibody, E74 like ETS transcription factor 1, ETS-related transcription factor Elf-1, ETS-like factor 1, EFTUD1, RIA1","show_quantity_input":1,"title":"ELF1 CUTANA™ CUT&RUN Antibody","gift_wrapping_available":false,"min_purchase_quantity":0,"customizations":[],"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/907/1009/chd3-cutana-cutandrun-antibody__66975.1645734525__91145.1648666822__83973.1648670498.jpg?c=2","alt":"ELF1 CUTANA™ CUT&RUN Antibody"}]} Pack Size: 100 µL
- Type: Polyclonal
- Host: Rabbit
- Applications: CUT&RUN, IHC
- Reactivity: Human
- Format: Antigen affinity-purified
- Target Size: 67 kDa
Description
This antibody meets EpiCypher’s "CUTANA Compatible" criteria for performance in Cleavage Under Targets and Release Using Nuclease (CUT&RUN) and/or Cleavage Under Targets and Tagmentation (CUT&Tag) approaches to genomic mapping. Every lot of a CUTANA Compatible antibody is tested in the indicated approach using EpiCypher optimized protocols and determined to yield peaks that show a genomic distribution pattern consistent with reported function(s) of the target protein. ELF1 is a member of the ETS (e26 transformation specific sequence) family of transcription factor proteins. ELF1 antibody produces CUT&RUN peaks above background primarily in intronic and promoter regions (Figure 1) that overlap with known ELF1 DNA-binding motifs (Figure 2). ELF1 is expressed in B and T cells and has been shown to be involved with the regulation of several T- and B-cell specific genes [1].
Validation Data
CUT&RUN was performed on 500k HeLa cells with 0.5 µg of either ELF1 or IgG negative control (EpiCypher 13-0042) antibodies using the CUTANA™ ChIC/CUT&RUN Kit v2.0 (EpiCypher 14-1048). Library preparation was performed with 5 ng of DNA (or the total amount recovered if less than 5 ng) using the CUTANA™ CUT&RUN Library Prep Kit (EpiCypher 14-1001/14-1002). Both kit protocols were adapted for high throughput Tecan liquid handling. Libraries were run on an Illumina NextSeq2000 with paired-end sequencing (2x50 bp). Sample sequencing depth was 5.0 million reads (IgG) and 6.0 million reads (ELF1). Data were aligned to the hg19 genome using Bowtie2. Data were filtered to remove duplicates, multi-aligned reads, and ENCODE DAC Exclusion List regions.
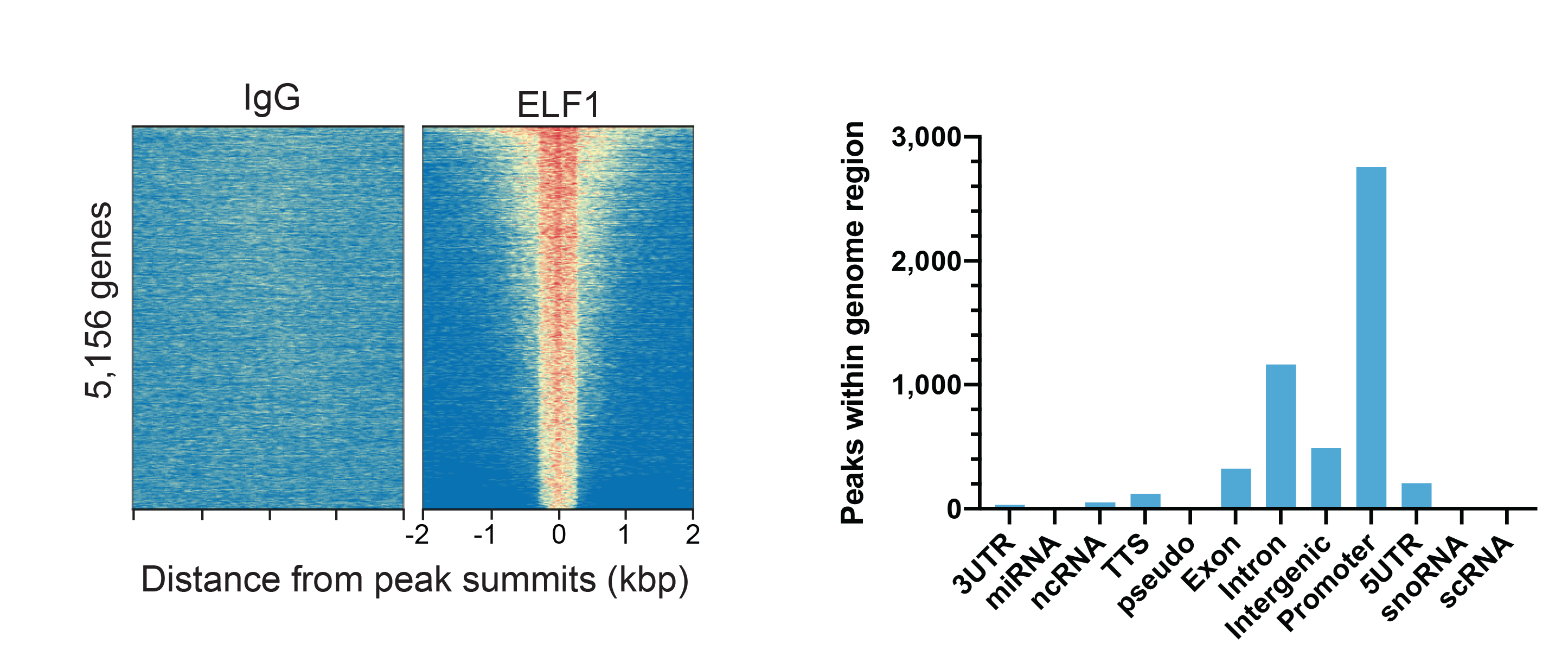
Figure 1: ELF1 peaks in CUT&RUN.
CUT&RUN was performed as described above. Peaks were called
using MACS2. Heatmaps show ELF1 peaks relative to IgG
negative control antibody in aligned rows ranked by
intensity (top to bottom) and color such that red indicates
high localized enrichment and blue denotes background signal
(left). The number of peaks that fall into
distinct classes of functionally annotated genomic regions
are shown (right).
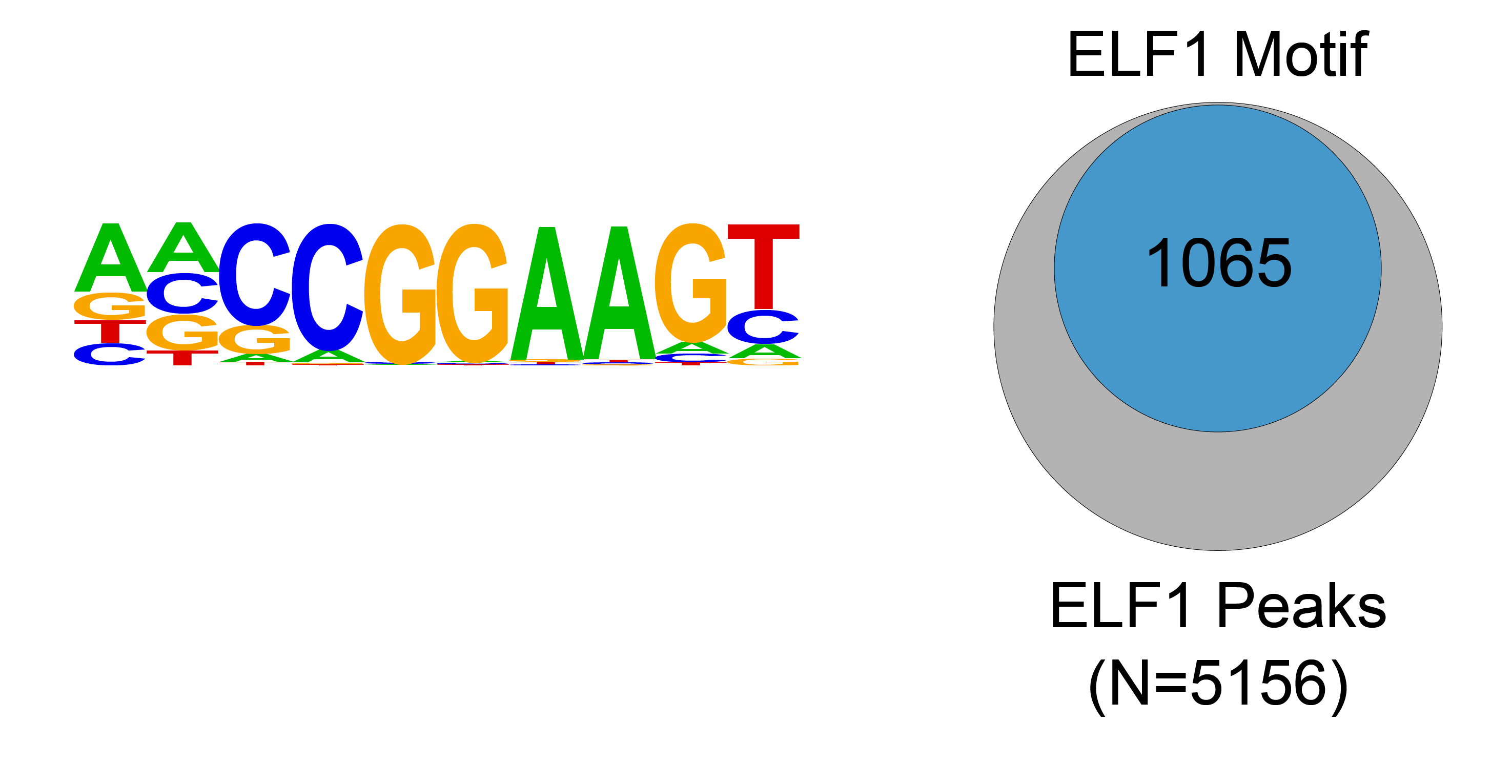
Figure 2: ELF1 transcription factor binding motif
analysis in CUT&RUN
Homer analysis determined that the ELF1(ETS)/Jurkat-ELF1-
ChIP-Seq(SRA014231)/Homer consensus motif, represented as a
sequence logo position weight matrix, was significantly
enriched under ELF1 CUT&RUN peaks (left).
The number of ELF1 peaks containing the consensus motif from
the left panel is represented by a Venn Diagram
(right).
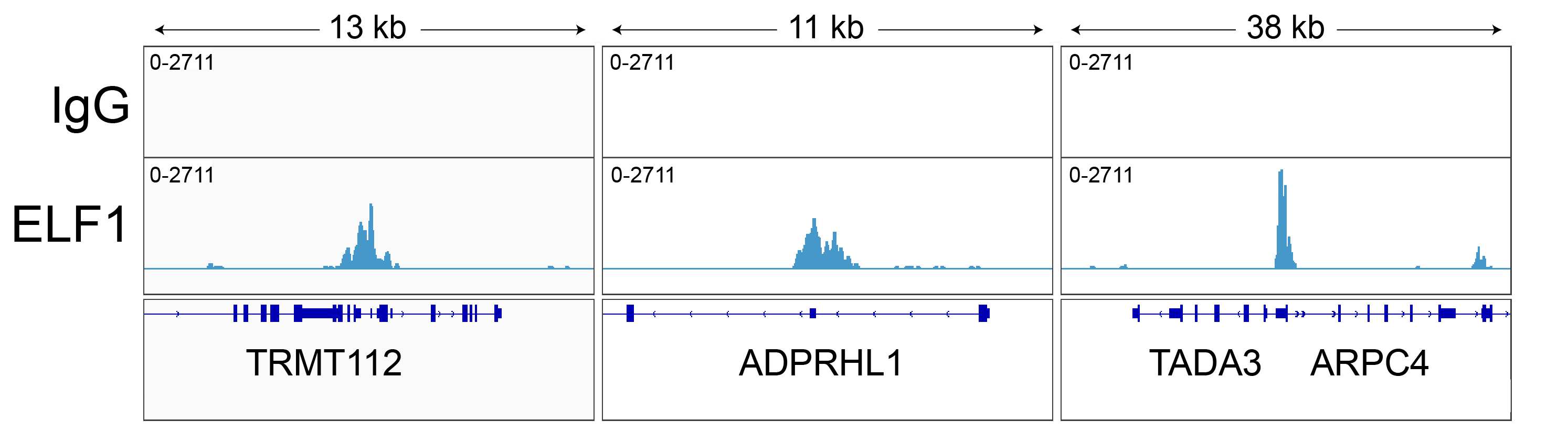
Figure 3: ELF1 CUT&RUN representative browser tracks
CUT&RUN was performed as described above. Gene browser shots
were generated using the Integrative Genomics Viewer (IGV,
Broad Institute). Three representative loci of the top
called peaks are shown.
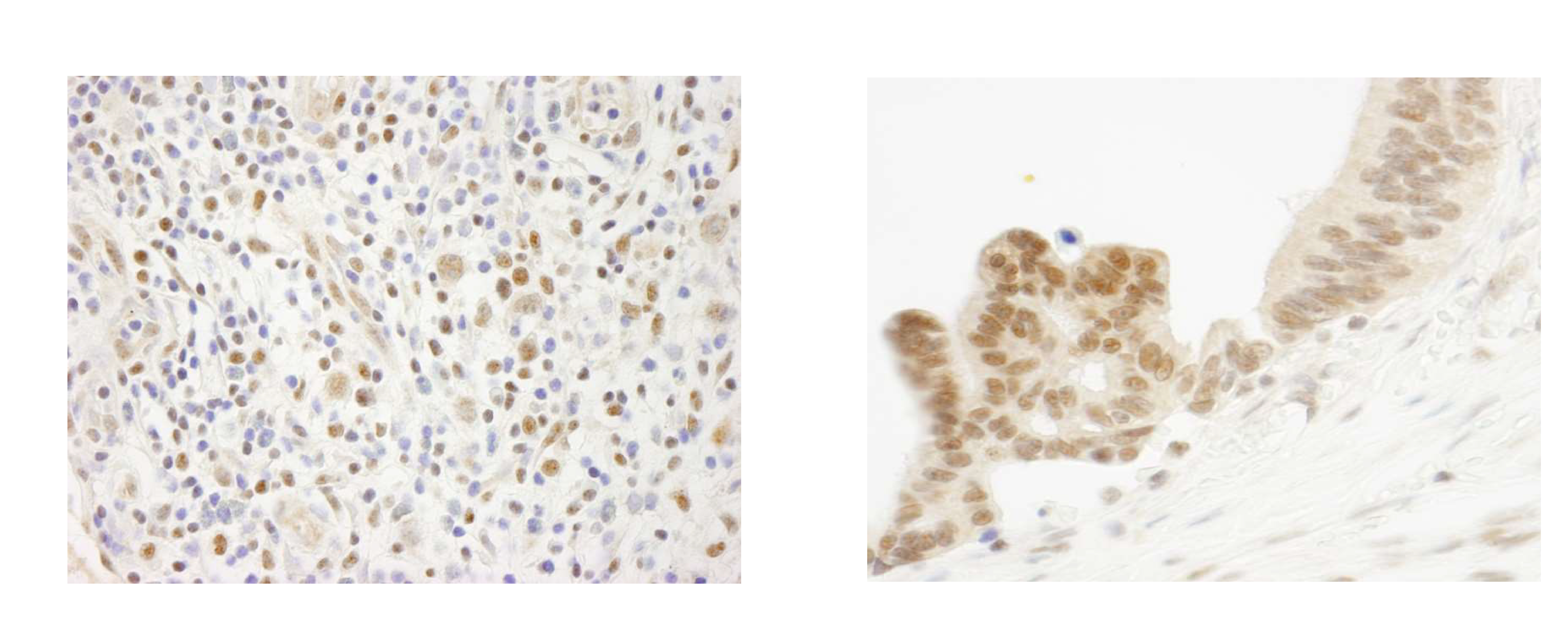
Figure 4: Immunohistochemistry data
FFPE sections of human Hodgkin’s Lymphoma
(left) and human ovarian carcinoma
(right) using ELF1 antibody at a dilution
of 1:250.
Technical Information
Recommended Dilution
Gene & Protein Information
References
Documents & Resources
Additional Info
This antibody is provided for commercial sale under license from Bethyl Laboratories, Inc.