Blog
March 25, 2026
Interrupting the epigenetic programs that drive tumor growth is a promising avenue for cancer therapy,...
February 24, 2026
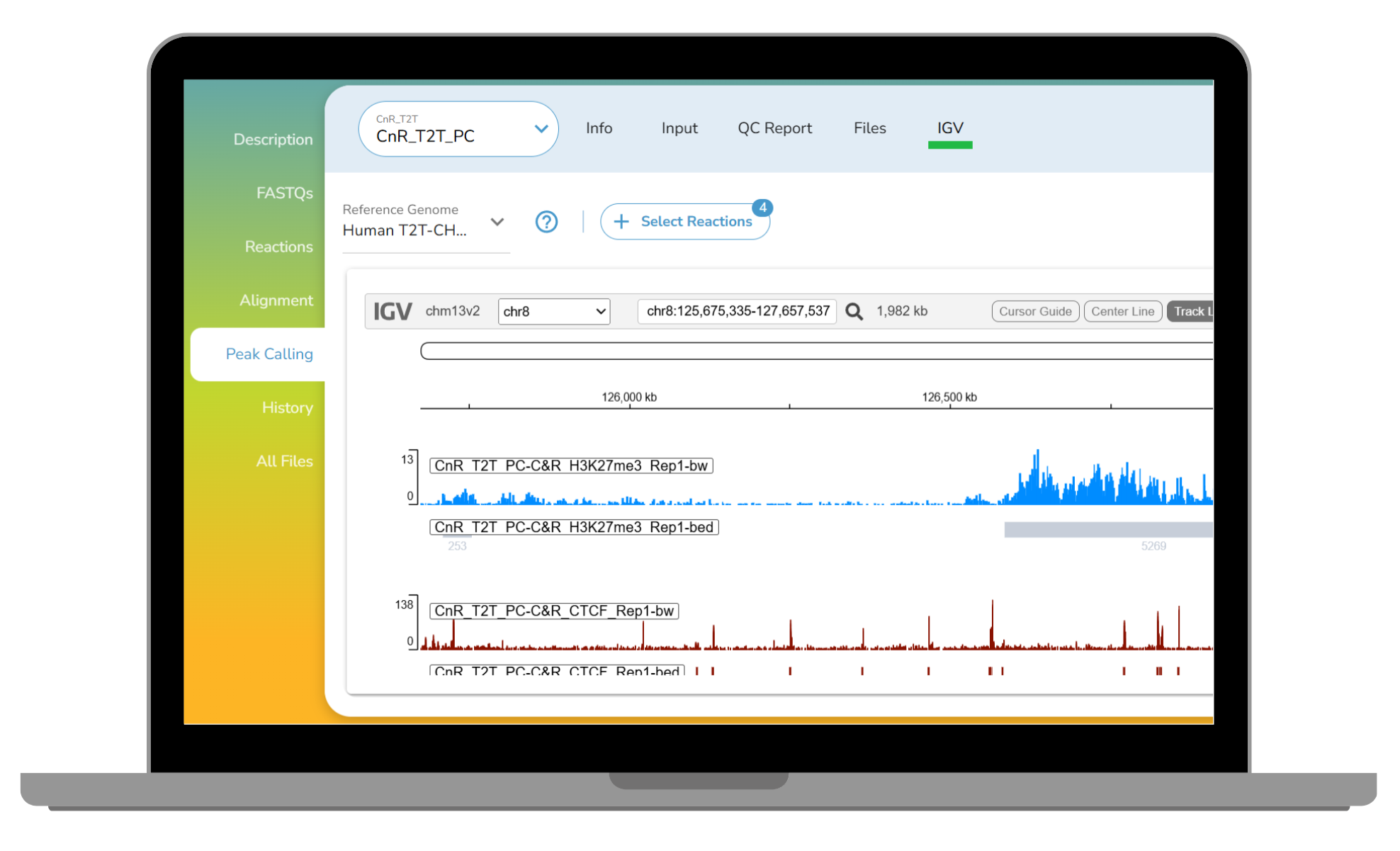
CUT&RUN and CUT&Tag have become gold-standard methods for profiling protein–DNA interactions,...

November 18, 2025
Durham, NC – Oct. 16, 2025 (The Scientist) – Winning a spot in the 2025 Top Innovations contest’s...

November 13, 2025
Epigenetic mechanisms shape nearly every aspect of neurobiology—from guiding early neural development...

October 28, 2025
Funding challenges are a reality for many researchers, and we want to make sure innovation doesn’t slow...

October 13, 2025
How do cells control when and where genes are expressed? The answer lies in a complex interplay of regulatory...

September 11, 2025
Gene expression is regulated by the complex molecular interplay between DNA methylation and chromatin...

September 3, 2025
Durham, NC – Sept. 3, 2025 (Life Science Newswire) – EpiCypher®, a pioneer in epigenomic...

June 12, 2025
Welcome to the new EpiCypher website experience! We did our best to design it with YOU, our Customer,...

June 11, 2025
Disruption of epigenetic mechanisms can lead to abnormal gene expression and is linked to many diseases,...

June 10, 2025
DNA methylation is a cornerstone of epigenetic regulation — and mapping it accurately across the genome...
No posts found