Nucleosome, Recombinant Human, Acidic Patch Mutant H2AE92K Biotinylated
{"url":"https://www.epicypher.com/products/nucleosomes/mutant-nucleosomes/nucleosome-recombinant-human-acidic-patch-mutant-h2ae92k-biotinylated","add_this":[{"service":"facebook","annotation":""},{"service":"email","annotation":""},{"service":"print","annotation":""},{"service":"twitter","annotation":""},{"service":"linkedin","annotation":""}],"gtin":null,"id":862,"bulk_discount_rates":[],"can_purchase":true,"meta_description":"Acidic patch mutant recombinant mononucleosomes (biotinylated H2AE92K), functionally validated with exceptional quality controls. Useful for chromatin binding, enzymatic, and structural studies.","category":["Nucleosomes/Mutant Nucleosomes","Nucleosomes/Mutant Nucleosomes/Acidic Patch Nucleosomes"],"AddThisServiceButtonMeta":"","main_image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/862/972/nucleosome-recombinant-human-acidic-patch-mutant-h2ae92k-biotinylated__90670.1645734564.jpg?c=2","alt":"Nucleosome, Recombinant Human, Acidic Patch Mutant H2AE92K Biotinylated"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=862","shipping":{"calculated":true},"num_reviews":0,"weight":"0.00 LBS","custom_fields":[{"id":"805","name":"Pack Size","value":"50 µg"}],"sku":"16-0030","description":"<div class=\"product-general-info\">\n <ul class=\"product-general-info__list-left\">\n <li class=\"product-general-info__list-item\">\n <strong>Species: </strong>Human\n </li>\n <li class=\"product-general-info__list-item\">\n <strong>Source: </strong><em>E. coli</em> & synthetic DNA\n </li>\n </ul>\n <ul class=\"product-general-info__list-right\">\n <li class=\"product-general-info__list-item\">\n <strong>Tag: </strong>Biotinylated\n </li>\n <li class=\"product-general-info__list-item\">\n <strong>Molecular Weight: </strong>199,741 Da\n </li>\n </ul>\n</div>\n<div class=\"service_accordion product-droppdown\">\n <div class=\"container\">\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel current\">\n <h3 class=\"sub-title1\">Description</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description-specific\"\n >\n <p>\n Recombinant mononucleosomes consist of 147 base pairs of DNA wrapped\n around an octamer core of histone proteins (two each of H2A, H2B,\n H3.1, and H4) to form a nucleosome, the basic repeating unit of\n chromatin. The 147 bp 601 sequence, which contains a 5’ biotin-TEG\n group, has high affinity for histone octamers and is useful for\n nucleosome assembly [1]. Histone H2A contains a glutamate-to-lysine\n (E-to-K) substitution at position 92 (H2AE92K). H2AE92 is among key\n residues forming a negatively charged region on the nucleosome\n surface named the “acidic patch”. The acidic patch is a conserved\n interaction hub for neighboring nucleosomes and nucleosome binding\n proteins, often via salt bridges with arginine anchors, and is\n functionally critical in chromatin condensation and chromatin\n remodeling [2-4]. H2AE92 resides in the H2A C-terminal extension and\n is associated with nucleosome binding factors such as histone H4\n N-terminal tail, LANA, RCC1, IL-33, SIR3, HMGN2, and remodeling\n ATPases [2-4]. H2AE92K disrupts binding with SMARCB1 [5].\n </p>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel current\">\n <h3 class=\"sub-title1\">Validation Data</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description-specific\"\n >\n <section class=\"image-picker\">\n <div class=\"image-picker__left\">\n <div\n class=\"image-picker__main-content_active image-picker__main-content\"\n >\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg\n class=\"image-picker__svg-left\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\"\n >\n <path\n d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\"\n />\n </svg>\n </button>\n <a\n href=\"/content/images/products/nucleosomes/16-0030-protein-gel-data-test.jpeg\"\n target=\"_blank\"\n class=\"image-picker__main-image-link\"\n ><img\n alt=\"16-0030-protein-gel-data-test\"\n src=\"/content/images/products/nucleosomes/16-0030-protein-gel-data-test.jpeg\"\n class=\"image-picker__main-image\"\n />\n <span class=\"image-picker__main-image-caption\"\n >(Click to enlarge)</span\n ></a\n >\n <button class=\"image-picker__right-arrow\">\n <svg\n class=\"image-picker__svg-right\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\"\n >\n <path\n d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\"\n />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\"\n ><strong>Figure 1: Protein gel data </strong><br />\n Coomassie stained PAGE gel of proteins in H2AE92K\n mononucleosome (1 µg) demonstrates the purity of the\n histones in the preparation. Sizes of molecular weight\n markers and positions of the core histones (H2AE92K, H2B, H3\n and H4) are indicated.\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg\n class=\"image-picker__svg-left\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\"\n >\n <path\n d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\"\n />\n </svg>\n </button>\n <a\n href=\"/content/images/products/nucleosomes/16-0030-dna-gel-data-test.jpeg\"\n target=\"_blank\"\n class=\"image-picker__main-image-link\"\n ><img\n alt=\"16-0030-dna-gel-data-test\"\n src=\"/content/images/products/nucleosomes/16-0030-dna-gel-data-test.jpeg\"\n class=\"image-picker__main-image\"\n />\n <span class=\"image-picker__main-image-caption\"\n >(Click to enlarge)</span\n ></a\n >\n <button class=\"image-picker__right-arrow\">\n <svg\n class=\"image-picker__svg-right\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\"\n >\n <path\n d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\"\n />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\"\n ><strong>Figure 2: DNA gel data </strong><br />\n H2AE92K mononucleosome resolved by native PAGE and stained\n with ethidium bromide to visualize DNA.\n <strong>Lane 1:</strong> Free DNA (EpiCypher\n <a\n href=\"/products/nucleosomes/nucleosome-assembly-601-sequence-dna-biotinylated\"\n >18-0005</a\n >; 100 ng). <strong>Lane 2:</strong> Intact H2AE92K\n mononucleosomes (400 ng).\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg\n class=\"image-picker__svg-left\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\"\n >\n <path\n d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\"\n />\n </svg>\n </button>\n <a\n href=\"/content/images/products/nucleosomes/16-0030-functional-binding-assay-test.jpeg\"\n target=\"_blank\"\n class=\"image-picker__main-image-link\"\n >\n <img\n alt=\"16-0030-functional-binding-assay-test\"\n src=\"/content/images/products/nucleosomes/16-0030-functional-binding-assay-test.jpeg\"\n class=\"image-picker__main-image\"\n />\n <span class=\"image-picker__main-image-caption\"\n >(Click to enlarge)</span\n >\n </a>\n <button class=\"image-picker__right-arrow\">\n <svg\n class=\"image-picker__svg-right\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\"\n >\n <path\n d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\"\n />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\">\n <strong>Figure 3: Functional binding assay </strong><br />\n The presence of acidic patch mutations disrupts LANA peptide\n binding to recombinant nucleosomes (WT control, EpiCypher\n <a\n href=\"/products/nucleosomes/mononucleosomes-recombinant-human-biotinylated\"\n >16-0006</a\n >; H2AE61A, EpiCypher\n <a\n href=\"/products/nucleosomes/acidic-patch-nucleosomes/nucleosome-recombinant-human-acidic-patch-mutant-h2ae61a-biotinylated\"\n >16-0029</a\n >; H2AE92K, EpiCypher 16-0030; H2BE105A,E113A, EpiCypher\n <a\n href=\"/products/nucleosomes/acidic-patch-nucleosomes/nucleosome-recombinant-human-acidic-patch-mutant-h2be105a-e113a-biotinylated\"\n >16-0031</a\n >). The binding of GST-tagged LANA peptide to biotinylated\n recombinant nucleosomes was assessed by AlphaLISA assay\n using Streptavidin Donor Beads and Glutathione Acceptor\n Beads (PerkinElmer). The presence of H2A acidic patch\n mutations completely blocks LANA binding, while H2B\n mutations cause a decrease in LANA binding affinity.\n </span>\n </p>\n </div>\n </div>\n <aside class=\"image-picker__right\">\n <div class=\"image-picker__gallery\">\n <img\n alt=\"16-0030-protein-gel-data-test\"\n src=\"/content/images/products/nucleosomes/16-0030-protein-gel-data-test.jpeg\"\n width=\"200\"\n class=\"image-picker__side-image image-picker__side-image_active\"\n role=\"button\"\n />\n <img\n alt=\"16-0030_dna_gel-data\"\n src=\"/content/images/products/nucleosomes/16-0030-dna-gel-data-test.jpeg\"\n class=\"image-picker__side-image\"\n role=\"button\"\n />\n <img\n alt=\"16-0030-functional-binding-assay-test\"\n src=\"/content/images/products/nucleosomes/16-0030-functional-binding-assay-test.jpeg\"\n class=\"image-picker__side-image\"\n role=\"button\"\n />\n </div>\n </aside>\n </section>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Technical Information</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\"\n >\n <div class=\"product-tech-info\">\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Storage</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n Stable for six months at -80°C from date of receipt. For best\n results, aliquot and avoid freeze/thaws.\n </div>\n </div>\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Formulation</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n 10 mM Tris-HCl pH 7.5, 25 mM NaCl, 1 mM EDTA, 2 mM DTT, 20%\n glycerol. (27.3 μg protein, 50 μg DNA + protein)\n </div>\n </div>\n </div>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Application Notes</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\"\n >\n <p>\n H2AE92K mononucleosome is highly purified and suitable for a variety\n of applications to test the effect of acidic patch mutation on\n enzymatic activity or chromatin binding. The biotinylated DNA\n enables affinity binding applications\n </p>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Gene & Protein Information</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\"\n >\n <div class=\"product-tech-info\">\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>UniProt ID</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n H2A - P04908 (alt. names: H2A type 1-B/E, H2A.2, H2A/a,\n H2A/m)<br />\n H2B - O60814 (alt. names: H2B K, HIRA-interacting protein 1)<br />\n H3.1 - P68431 (alt. names: H3, H3/a, H3/b, H3/c, H3/d)<br />\n H4 - P62805\n </div>\n </div>\n </div>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">References</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\"\n >\n <strong>Background References:</strong>\n <br />\n [1] Lowary & Widom <em>J. Mol. Biol.</em> (1998). PMID:\n <a\n href=\"https://pubmed.ncbi.nlm.nih.gov/9514715/\"\n target=\"_blank\"\n title=\"New DNA sequence rules for high affinity binding to histone octamer and sequence-directed nucleosome positioning\"\n >9514715</a\n ><br />\n [2] Kalashnikova et al. <em>J. R. Soc. Interface</em> (2013). PMID:\n <a\n href=\"https://pubmed.ncbi.nlm.nih.gov/23446052/\"\n target=\"_blank\"\n title=\"The role of the nucleosome acidic patch in modulating higher order chromatin structure\"\n >23446052</a\n ><br />\n [3] Levendosky & Bowman <em>eLife</em> (2019). PMID:\n <a\n href=\"https://pubmed.ncbi.nlm.nih.gov/31094676/\"\n target=\"_blank\"\n title=\"Asymmetry between the two acidic patches dictates the direction of nucleosome sliding by the ISWI chromatin remodeler\"\n >31094676</a\n ><br />\n [4] Gamarra et al. <em>eLife</em> (2018). PMID:\n <a\n href=\"https://pubmed.ncbi.nlm.nih.gov/29664398/\"\n target=\"_blank\"\n title=\"The nucleosomal acidic patch relieves auto-inhibition by the ISWI remodeler SNF2h\"\n >29664398</a\n ><br />\n [5] Valencia et al. <em>Cell</em> (2019). PMID:\n <a\n href=\"https://pubmed.ncbi.nlm.nih.gov/31759698/\"\n target=\"_blank\"\n title=\"Recurrent SMARCB1 Mutations Reveal a Nucleosome Acidic Patch Interaction Site That Potentiates mSWI/SNF Complex Chromatin Remodeling\"\n >31759698</a\n ><br />\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Documents & Resources</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\"\n >\n <div class=\"product-documents\">\n <a\n href=\"/content/documents/tds/16-0030.pdf\"\n target=\"_blank\"\n class=\"product-documents__link\"\n >\n <svg\n version=\"1.1\"\n id=\"Layer_1\"\n xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\"\n x=\"0px\"\n y=\"0px\"\n viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\"\n xml:space=\"preserve\"\n class=\"product-documents__icon\"\n alt=\"16-0030 Datasheet\"\n >\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M191.92,68.77l-47.69-47.69c-1.33-1.33-3.12-2.08-5.01-2.08H45.09C41.17,19,38,22.17,38,26.09v184.36\n c0,3.92,3.17,7.09,7.09,7.09h141.82c3.92,0,7.09-3.17,7.09-7.09V73.8C194,71.92,193.25,70.1,191.92,68.77z M177.65,77.06h-41.7\n v-41.7L177.65,77.06z M178.05,201.59H53.95V34.95h66.92v47.86c0,5.14,4.17,9.31,9.31,9.31h47.86V201.59z\"\n />\n </g>\n <rect\n x=\"20\"\n y=\"112\"\n class=\"product-documents__svg-background\"\n width=\"146\"\n height=\"76\"\n />\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M23.83,125.68h22.36c5.29,0,9.41,1.33,12.35,4c2.94,2.67,4.42,6.39,4.42,11.18c0,4.78-1.47,8.51-4.42,11.18\n c-2.94,2.67-7.06,4-12.35,4H34.59v18.29H23.83V125.68z M44.81,147.9c5.38,0,8.07-2.32,8.07-6.97c0-2.39-0.67-4.16-2-5.31\n c-1.33-1.15-3.36-1.73-6.07-1.73H34.59v14.01H44.81z\"\n />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M69.92,125.68h18.91c5.29,0,9.84,0.97,13.66,2.9c3.82,1.93,6.74,4.72,8.76,8.35\n c2.02,3.63,3.04,7.98,3.04,13.04c0,5.06-1,9.42-3,13.08c-2,3.66-4.91,6.45-8.73,8.38c-3.82,1.93-8.4,2.9-13.73,2.9H69.92V125.68z\n M88.07,165.63c10.35,0,15.52-5.22,15.52-15.66c0-10.4-5.17-15.59-15.52-15.59h-7.38v31.26H88.07z\"\n />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M122.57,125.68h32.84v8.49h-22.22v11.18h20.84v8.49h-20.84v20.49h-10.63V125.68z\"\n />\n </g>\n </svg>\n <span class=\"product-documents__info\">Technical Datasheet</span>\n </a>\n </div>\n </div>\n </div>\n </div>\n </div>\n</div>\n","tags":[],"warranty":"","price":{"without_tax":{"formatted":"$475.00","value":475,"currency":"USD"},"tax_label":"Sales Tax"},"detail_messages":"","availability":"","page_title":"Acidic Patch Mutant Recombinant Nucleosome | H2AE92K, Biotinylated","cart_url":"https://www.epicypher.com/cart.php","max_purchase_quantity":0,"mpn":null,"upc":null,"options":[],"related_products":[{"id":865,"sku":"16-1030","name":"Nucleosome, Recombinant Human, Acidic Patch Mutant H2AE92K","url":"https://www.epicypher.com/products/nucleosomes/mutant-nucleosomes/nucleosome-recombinant-human-acidic-patch-mutant-h2ae92k","availability":"","rating":null,"brand":{"name":null},"category":["Nucleosomes/Mutant Nucleosomes","Nucleosomes/Mutant Nucleosomes/Acidic Patch Nucleosomes"],"summary":"\n \n \n \n Description\n \n \n Mono","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/865/890/2021.05.05_acidic_patch_nuc_RGB_white_border_LS_icon_thumbnail_board_2__87356.1626114896.png?c=2","alt":"Nucleosome, Recombinant Human, Acidic Patch Mutant H2AE92K"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/865/890/2021.05.05_acidic_patch_nuc_RGB_white_border_LS_icon_thumbnail_board_2__87356.1626114896.png?c=2","alt":"Nucleosome, Recombinant Human, Acidic Patch Mutant H2AE92K"}],"date_added":"23rd May 2021","pre_order":false,"show_cart_action":true,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":802,"name":"Pack Size","value":"50 µg"}],"num_reviews":null,"weight":{"formatted":"0.00 LBS","value":0},"demo":false,"add_to_cart_url":"https://www.epicypher.com/cart.php?action=add&product_id=865","price":{"without_tax":{"currency":"USD","formatted":"$475.00","value":475},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=865"},{"id":705,"sku":"16-2030","name":"Acidic Patch Mutant H2AE92K Recombinant Nucleosome with Linker DNA, Biotinylated","url":"https://www.epicypher.com/products/nucleosomes/mutant-nucleosomes/acidic-patch-mutant-h2ae92k-recombinant-nucleosome-with-linker-dna-biotinylated","availability":"","rating":null,"brand":{"name":null},"category":["Nucleosomes","Nucleosomes/Mutant Nucleosomes","Nucleosomes/Mutant Nucleosomes/Acidic Patch Nucleosomes"],"summary":"\n \n \n Species: Human\n \n \n Source: E. coli & synthetic DNA\n \n \n \n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/705/1227/2021.05.05_acidic_patch_nuc_RGB_white_border_LS_icon_thumbnail_board_2__41959__47696.1717694541.png?c=2","alt":"Acidic Patch Mutant H2AE92K Recombinant Nucleosome with Linker DNA, Biotinylated"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/705/1227/2021.05.05_acidic_patch_nuc_RGB_white_border_LS_icon_thumbnail_board_2__41959__47696.1717694541.png?c=2","alt":"Acidic Patch Mutant H2AE92K Recombinant Nucleosome with Linker DNA, Biotinylated"}],"date_added":"18th Oct 2019","pre_order":false,"show_cart_action":true,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":492,"name":"Pack size","value":"50 µg"}],"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"add_to_cart_url":"https://www.epicypher.com/cart.php?action=add&product_id=705","price":{"without_tax":{"currency":"USD","formatted":"$475.00","value":475},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=705"},{"id":863,"sku":"16-0029","name":"Nucleosome, Recombinant Human, Acidic Patch Mutant H2AE61A Biotinylated","url":"https://www.epicypher.com/products/nucleosomes/mutant-nucleosomes/nucleosome-recombinant-human-acidic-patch-mutant-h2ae61a-biotinylated","availability":"","rating":null,"brand":{"name":null},"category":["Nucleosomes/Mutant Nucleosomes","Nucleosomes/Mutant Nucleosomes/Acidic Patch Nucleosomes"],"summary":"\n \n \n Species: Human\n \n \n Source: E. coli & synthetic DNA\n \n \n \n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/863/888/2021.05.05_acidic_patch_nuc_RGB_white_border_LS_icon_thumbnail_board_2__67572.1621809858.png?c=2","alt":"Nucleosome, Recombinant Human, Acidic Patch Mutant H2AE61A Biotinylated"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/863/888/2021.05.05_acidic_patch_nuc_RGB_white_border_LS_icon_thumbnail_board_2__67572.1621809858.png?c=2","alt":"Nucleosome, Recombinant Human, Acidic Patch Mutant H2AE61A Biotinylated"}],"date_added":"23rd May 2021","pre_order":false,"show_cart_action":true,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":804,"name":"Pack Size","value":"50 µg"}],"num_reviews":null,"weight":{"formatted":"0.00 LBS","value":0},"demo":false,"add_to_cart_url":"https://www.epicypher.com/cart.php?action=add&product_id=863","price":{"without_tax":{"currency":"USD","formatted":"$475.00","value":475},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=863"},{"id":861,"sku":"16-0031","name":"Nucleosome, Recombinant Human, Acidic Patch Mutant H2BE105A,E113A Biotinylated","url":"https://www.epicypher.com/products/nucleosomes/mutant-nucleosomes/nucleosome-recombinant-human-acidic-patch-mutant-h2be105a-e113a-biotinylated","availability":"","rating":null,"brand":{"name":null},"category":["Nucleosomes/Mutant Nucleosomes","Nucleosomes/Mutant Nucleosomes/Acidic Patch Nucleosomes"],"summary":"\n \n \n Species: Human\n \n \n Source: E. coli & synthetic DNA\n \n \n \n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/861/927/nucleosome-recombinant-human-acidic-patch-mutant-h2be105ae113a-biotinylated__42717.1645734518.jpg?c=2","alt":"Nucleosome, Recombinant Human, Acidic Patch Mutant H2BE105A,E113A Biotinylated"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/861/927/nucleosome-recombinant-human-acidic-patch-mutant-h2be105ae113a-biotinylated__42717.1645734518.jpg?c=2","alt":"Nucleosome, Recombinant Human, Acidic Patch Mutant H2BE105A,E113A Biotinylated"},{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/861/885/2021.05.05_acidic_patch_nuc_RGB_LS__97220.1621279281.png?c=2","alt":"Nucleosome, Recombinant Human, Acidic Patch Mutant H2BE105A,E113A Biotinylated"}],"date_added":"17th May 2021","pre_order":false,"show_cart_action":true,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":806,"name":"Pack Size","value":"50 µg"}],"num_reviews":null,"weight":{"formatted":"0.00 LBS","value":0},"demo":false,"add_to_cart_url":"https://www.epicypher.com/cart.php?action=add&product_id=861","price":{"without_tax":{"currency":"USD","formatted":"$475.00","value":475},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=861"},{"id":864,"sku":"16-1029","name":"Nucleosome, Recombinant Human, Acidic Patch Mutant H2AE61A","url":"https://www.epicypher.com/products/nucleosomes/mutant-nucleosomes/nucleosome-recombinant-human-acidic-patch-mutant-h2ae61a","availability":"","rating":null,"brand":{"name":null},"category":["Nucleosomes/Mutant Nucleosomes","Nucleosomes/Mutant Nucleosomes/Acidic Patch Nucleosomes"],"summary":"\n \n \n \n Description\n \n \n Mono","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/864/961/nucleosome-recombinant-human-acidic-patch-mutant-h2ae61a__21865.1645734553.jpg?c=2","alt":"Nucleosome, Recombinant Human, Acidic Patch Mutant H2AE61A"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/864/961/nucleosome-recombinant-human-acidic-patch-mutant-h2ae61a__21865.1645734553.jpg?c=2","alt":"Nucleosome, Recombinant Human, Acidic Patch Mutant H2AE61A"}],"date_added":"23rd May 2021","pre_order":false,"show_cart_action":true,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":803,"name":"Pack Size","value":"50 µg"}],"num_reviews":null,"weight":{"formatted":"0.00 LBS","value":0},"demo":false,"add_to_cart_url":"https://www.epicypher.com/cart.php?action=add&product_id=864","price":{"without_tax":{"currency":"USD","formatted":"$475.00","value":475},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=864"}],"shipping_messages":[],"rating":0,"meta_keywords":"acidic patch mutant, mutant nucleosome, mutant histone, H2AE92K","show_quantity_input":1,"title":"Nucleosome, Recombinant Human, Acidic Patch Mutant H2AE92K Biotinylated","gift_wrapping_available":false,"min_purchase_quantity":0,"customizations":[],"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/862/972/nucleosome-recombinant-human-acidic-patch-mutant-h2ae92k-biotinylated__90670.1645734564.jpg?c=2","alt":"Nucleosome, Recombinant Human, Acidic Patch Mutant H2AE92K Biotinylated"}]} Pack Size: 50 µg
- Species: Human
- Source: E. coli & synthetic DNA
- Tag: Biotinylated
- Molecular Weight: 199,741 Da
Description
Recombinant mononucleosomes consist of 147 base pairs of DNA wrapped around an octamer core of histone proteins (two each of H2A, H2B, H3.1, and H4) to form a nucleosome, the basic repeating unit of chromatin. The 147 bp 601 sequence, which contains a 5’ biotin-TEG group, has high affinity for histone octamers and is useful for nucleosome assembly [1]. Histone H2A contains a glutamate-to-lysine (E-to-K) substitution at position 92 (H2AE92K). H2AE92 is among key residues forming a negatively charged region on the nucleosome surface named the “acidic patch”. The acidic patch is a conserved interaction hub for neighboring nucleosomes and nucleosome binding proteins, often via salt bridges with arginine anchors, and is functionally critical in chromatin condensation and chromatin remodeling [2-4]. H2AE92 resides in the H2A C-terminal extension and is associated with nucleosome binding factors such as histone H4 N-terminal tail, LANA, RCC1, IL-33, SIR3, HMGN2, and remodeling ATPases [2-4]. H2AE92K disrupts binding with SMARCB1 [5].
Validation Data
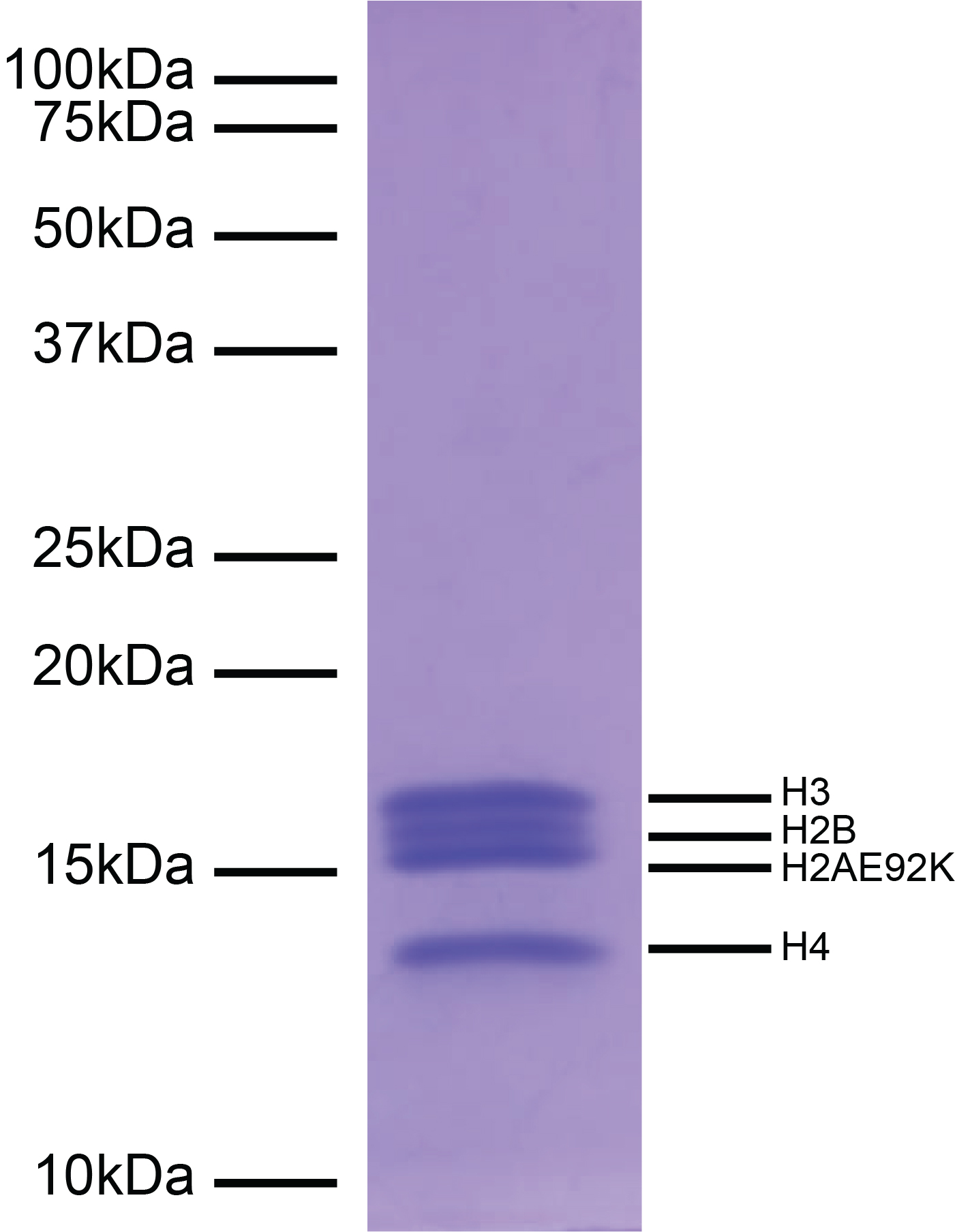
Figure 1: Protein gel data
Coomassie stained PAGE gel of proteins in H2AE92K
mononucleosome (1 µg) demonstrates the purity of the
histones in the preparation. Sizes of molecular weight
markers and positions of the core histones (H2AE92K, H2B, H3
and H4) are indicated.
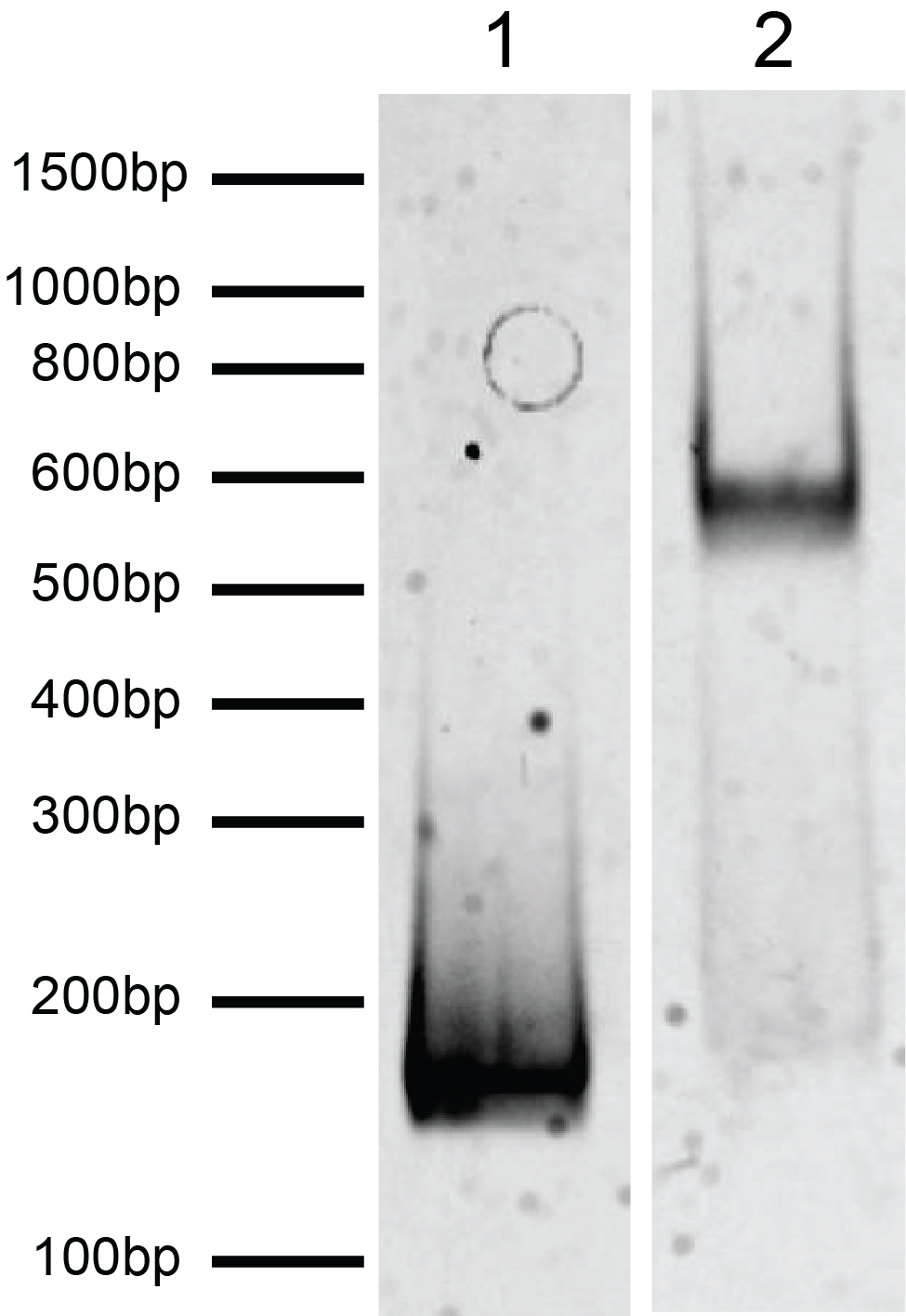
Figure 2: DNA gel data
H2AE92K mononucleosome resolved by native PAGE and stained
with ethidium bromide to visualize DNA.
Lane 1: Free DNA (EpiCypher
18-0005; 100 ng). Lane 2: Intact H2AE92K
mononucleosomes (400 ng).
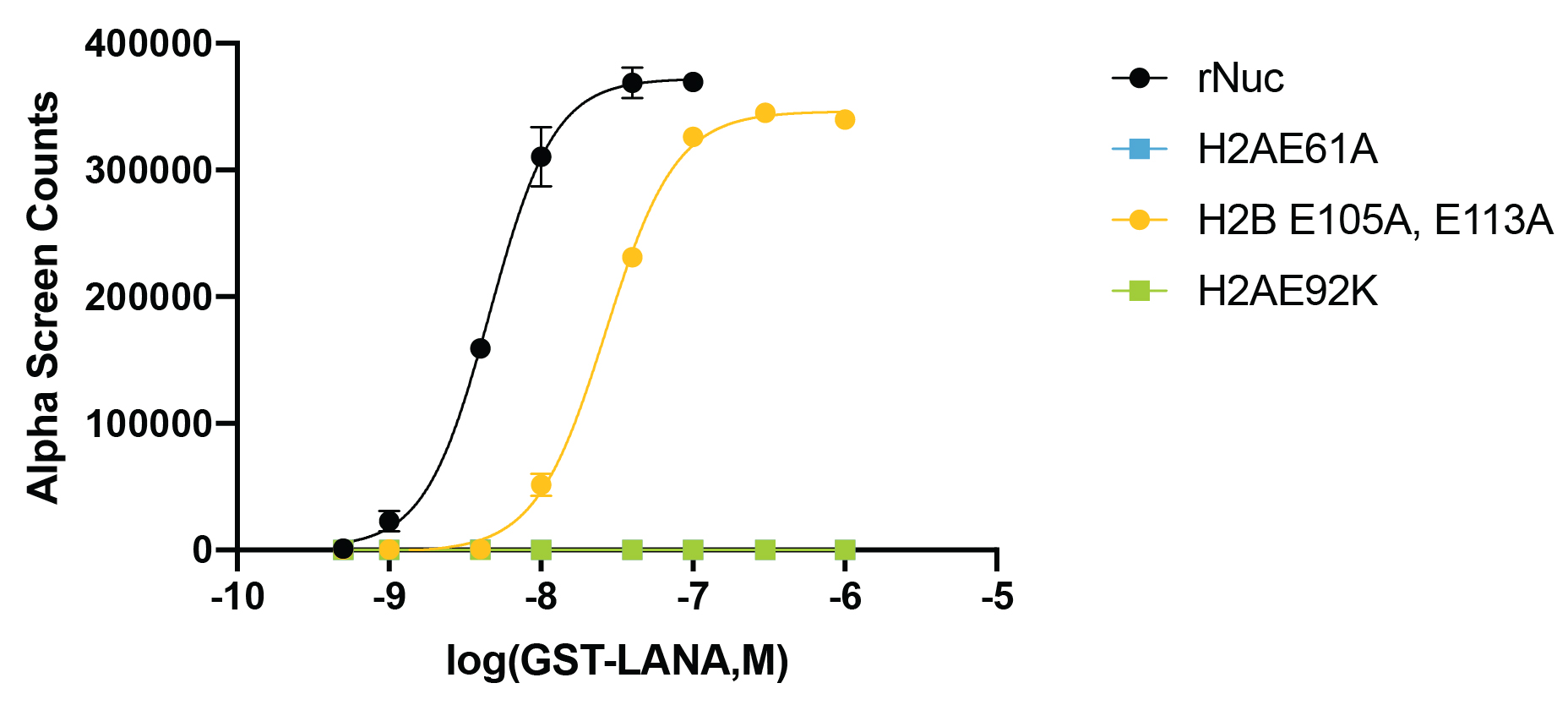
Figure 3: Functional binding assay
The presence of acidic patch mutations disrupts LANA peptide
binding to recombinant nucleosomes (WT control, EpiCypher
16-0006; H2AE61A, EpiCypher
16-0029; H2AE92K, EpiCypher 16-0030; H2BE105A,E113A, EpiCypher
16-0031). The binding of GST-tagged LANA peptide to biotinylated
recombinant nucleosomes was assessed by AlphaLISA assay
using Streptavidin Donor Beads and Glutathione Acceptor
Beads (PerkinElmer). The presence of H2A acidic patch
mutations completely blocks LANA binding, while H2B
mutations cause a decrease in LANA binding affinity.
Technical Information
Application Notes
H2AE92K mononucleosome is highly purified and suitable for a variety of applications to test the effect of acidic patch mutation on enzymatic activity or chromatin binding. The biotinylated DNA enables affinity binding applications
Gene & Protein Information
H2B - O60814 (alt. names: H2B K, HIRA-interacting protein 1)
H3.1 - P68431 (alt. names: H3, H3/a, H3/b, H3/c, H3/d)
H4 - P62805