H3K4me3 Antibody, SNAP-Certified™ for CUT&RUN – DISCONTINUED
Out of stock
Product Discontinued – see 13-0060 as the recommended alternative
This H3K4me3 (histone H3 lysine 4 trimethyl) antibody meets EpiCypher’s lot-specific SNAP-Certified™ criteria for specificity and efficient target enrichment in CUT&RUN. This requires <20% cross-reactivity to related histone PTMs determined using the SNAP-CUTANA™ K-MetStat Panel of spike-in controls (EpiCypher 19-1002, Figure 1). High target efficiency is confirmed by consistent genomic enrichment at 500k and 50k starting cells (Figures 2-4). This antibody targets histone H3 trimethylated at lysine 4, which is enriched at active promoters near transcription start sites (TSS).
*Mixed Monoclonal: a pool of multiple recombinant monoclonal antibodies.
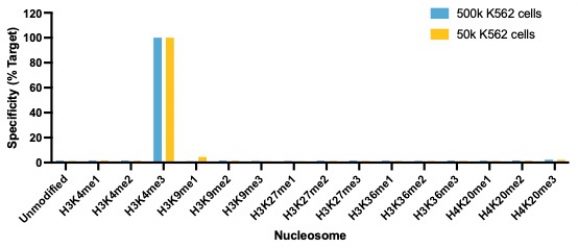
Figure 1: SNAP specificity analysis in CUT&RUN
Figure 2: CUT&RUN genome wide enrichment
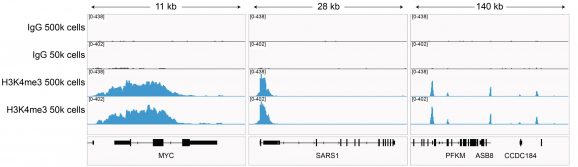
Figure 3: H3K4me3 CUT&RUN representative browser tracks
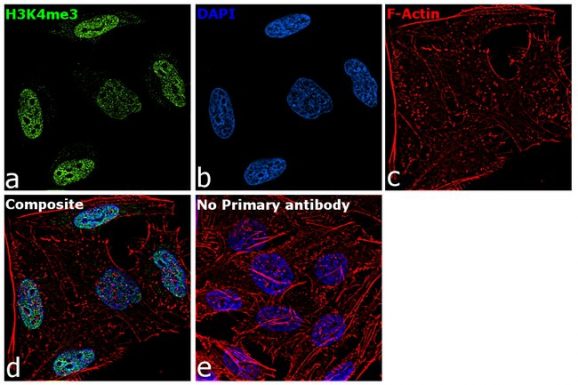
Figure 4: Immunofluorescence
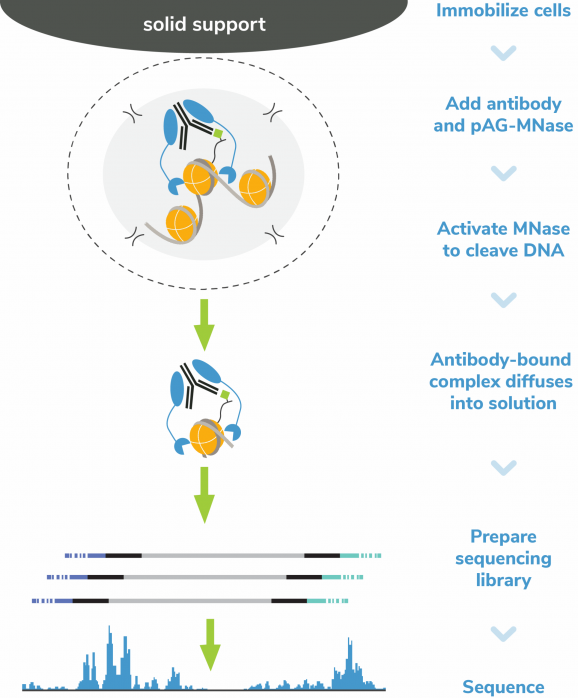
Figure 6: CUT&RUN methods
CUT&RUN was performed on 500k and 50k K562 cells with the SNAP-CUTANA™ K-MetStat Panel (EpiCypher 19-1002) spiked-in prior to the addition of 0.5 μg of either H3K4me3 or IgG negative control (EpiCypher 13-0042) antibodies. The experiment was performed using the CUTANA™ ChIC/CUT&RUN Kit v3 (EpiCypher 14-1048). Library preparation was performed with 5 ng of CUT&RUN enriched DNA (or the total amount recovered if less than 5 ng) using the CUTANA™ CUT&RUN Library Prep Kit (EpiCypher 14-1001/14-1002). Both kit protocols were adapted for high throughput Tecan liquid handling. Libraries were run on an Illumina NextSeq2000 with paired-end sequencing (2×50 bp). Reaction sequencing depth was 8.3 million reads (IgG 500k cell input), 15.5 million reads (IgG 50k cell input), 9.8 million reads (H3K4me3 500k cell input) and 8.6 million reads (H3K4me3 50k cell input). Data were aligned to the hg19 genome using Bowtie2. Data were filtered to remove duplicates, multi-aligned reads, and ENCODE DAC Exclusion List regions.
- Type: Mixed Monoclonal*
- Host: Rabbit
- Applications: CUT&RUN, ICC/IF, WB
NOTE: Previous lots were validated in ChIP, however lots are no longer tested in this application.
- Reactivity: Human, Mouse, Drosophila, Yeast, Wide Range (Predicted)
- Format: Protein A affinity-purified
- Target Size: 15 kDa
- CUT&RUN: 0.5 µg per reaction
- Immunofluorescence (IF): 1:100
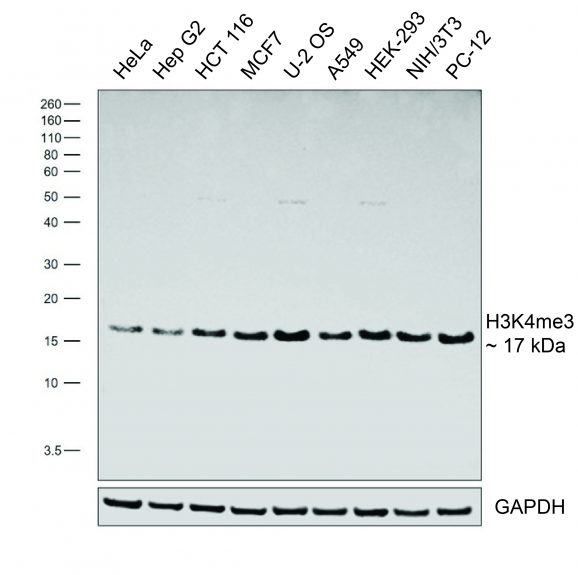
- Western Blot (WB): 1:250