Putting CUT&RUN/Tag Data Analysis Back in the Hands of Biologists
- Nathan Hall
CUT&RUN and CUT&Tag have become gold-standard methods for profiling protein–DNA interactions, offering high resolution, low background, and efficient sequencing. While these assays dramatically simplified experimental workflows, data analysis has remained a major barrier. For many labs, generating sequencing data is no longer the hard part — turning FASTQs into confident biological conclusions is.
CUTANA™ Cloud was created to close that gap, delivering validated, accessible CUT&RUN and CUT&Tag data analysis directly to the scientists generating the data.
The Historical Challenge: Powerful Assays, Limited Access to Analysis
Despite the experimental advantages of these assays, CUT&RUN and CUT&Tag data analysis still relies on a fragmented ecosystem of command-line tools, custom scripts, and lab-specific workflows. Alignment, quality control, normalization, peak calling, and visualization are often stitched together differently in every lab, with little standardization and limited transparency.
For many researchers, this has meant:
- Dependence on bioinformaticians or core facilities
- Long turnaround times between sequencing and insight
- Difficulty interpreting QC metrics or troubleshooting experiments
- Limited confidence that pipelines are configured correctly
The result is a persistent disconnect between experimental success and analytical certainty.
Built on Proven Science, Not Theoretical Pipelines
CUTANA Cloud is powered by the same analysis pipelines EpiCypher has used internally to analyze over 30,000 CUT&RUN and CUT&Tag reactions, resulting in 12 peer-reviewed studies (5 additional studies in preparation).
These pipelines were not assembled to be maximally flexible, but were refined through years of experimental iteration, benchmarking, and real-world use. That experience is embedded directly into CUTANA Cloud, ensuring every analysis reflects best practices for epigenomic profiling rather than generic NGS assumptions.
A Guided Workflow from FASTQ to Insight
CUTANA Cloud’s design is focused on the practical progression of CUT&RUN/Tag experiments, rather than being organized around abstract bioinformatics concepts and command line interfaces, the platform offers a simple user interface and easy to interpret results, providing utility to novices and experts alike.

Once data enters the platform, users move through a clear, linear workflow (Figure 2):

- Upload FASTQs directly or import from BaseSpaceTM
- Annotate reactions with experimental metadata and review FASTQC reports
- Align reads to supported reference genomes (human, mouse, drosophila, yeast)
- Review Alignment QC, including sequencing metrics and spike-in performance
- Call peaks using MACS2 or SICER2 with assay-appropriate defaults
- Explore results via an on-platform IGV browser
- Download standard files for downstream or local analysis
At every stage, outputs are immediately interpretable, not buried behind logs or scripts allowing the user to have a transparent understanding of the quality of their experiment without the need of digging through one tool’s output in relation to another.
Not a Black Box, Not a Bioinformatics Obstacle Course
Many existing platforms fall at one of two extremes: fully opaque pipelines that hide every decision, or hyper-customizable environments that expose dozens of parameters without guidance. CUTANA Cloud deliberately sits in between. Only parameters that meaningfully impact CUT&RUN and CUT&Tag outcomes are exposed. Everything else is fixed to validated defaults. This approach:
- Reduces cognitive overhead for new users
- Prevents misconfigured analyses
- Ensures datasets are cross-comparable across experiments and labs
- Preserves transparency without overwhelming complexity
This balance allows wet-lab scientists to run analyses confidently while also enabling bioinformatics-savvy users to move quickly through repetitive steps without sacrificing rigor.
Transparent QC, Built for Interpretation

Quality control is not treated as an afterthought. Each alignment generates a comprehensive QC report (Figure 3) that includes sequencing statistics, alignment metrics, and SNAP-CUTANA spike-in performance. These reports are designed to answer practical questions: Did the experiment work? Is enrichment specific? Is the dataset worth moving forward?
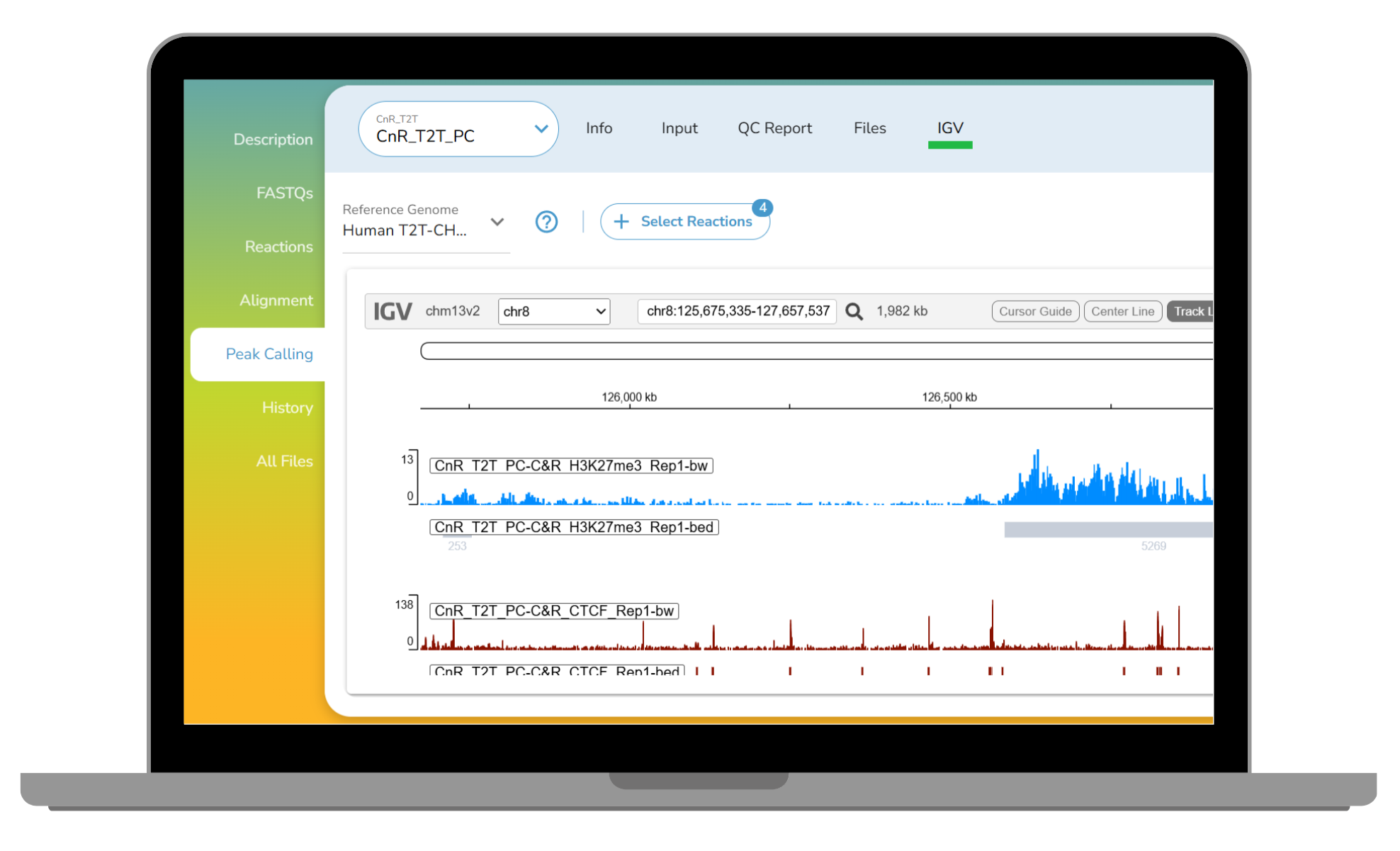
For deeper inspection, CUTANA Cloud includes an integrated IGV browser (Figure 4), allowing users to visually assess signal, peaks, and controls without exporting files or configuring local environments.

All outputs e.g. BAMs, bigWigs, BED files, FRiP scores, and peak annotations are standard and fully portable.
Designed for How Labs Actually Operate
CUTANA Cloud supports multiple user types without forcing a single workflow :
- Wet-lab scientists validating experimental success and exploring results in the absence of coding knowhow
- Core facilities running standardized, reproducible analyses at scale
- Bioinformaticians offloading alignment and QC to focus on higher-order analysis
Importantly, CUTANA Cloud uses a pay-as-you-go credit model (Figure 5). Labs are charged only for alignment runs; peak calling is included at no additional cost. There are no subscriptions or contracts, making the platform viable for groups running a handful of experiments per year as well as high- throughput power users. This pricing model reflects a simple reality: not every lab needs a five-figure annual license to analyze 20 reactions. CUTANA Cloud requires no subscription and no upfront commitment. CUTANA Cloud offers validated analysis, on demand.

Get Started with CUTANA Cloud
By embedding experimental expertise directly into accessible, validated pipelines, CUTANA Cloud removes the traditional handoff between experiment and analysis, moving the user from dependency to ownership. Scientists no longer need to wait for interpretation or rely on opaque workflows. Instead, they gain direct control over their data empowering faster feedback and greater confidence in downstream conclusions of their experiment.
To see the platform in action, create a free CUTANA Cloud account and explore the CUTANATM Gold Standard CUT&RUN and CUT&Tag datasets. These reference datasets let users interact with real QC reports, genome browser tracks, and outputs before uploading their own data.
Create an account and explore the Gold Standard datasets today.
