EpiCypher’s Optimized CUTANA™ CUT&RUN Protocol for Transcription Factors, Histone PTMs, and More!
- Ellen Weinzapfel
Immunotethering approaches, such as those used in CUT&RUN (Cleavage Under Targets & Release Using Nuclease) assays, are one of the most exciting advancements in chromatin biology, as they enable you to generate high-resolution chromatin profiling data in a fraction of the time and cost of standard ChIP-seq (Chromatin Immunoprecipitation-sequencing) experiments1-3. EpiCypher is leveraging these technological innovations to deliver transformative tools for epigenetics research, starting with the launch of CUTANATM pAG-MNase for ChIC / CUT&RUN assays.
How do CUTANATM CUT&RUN assays work?
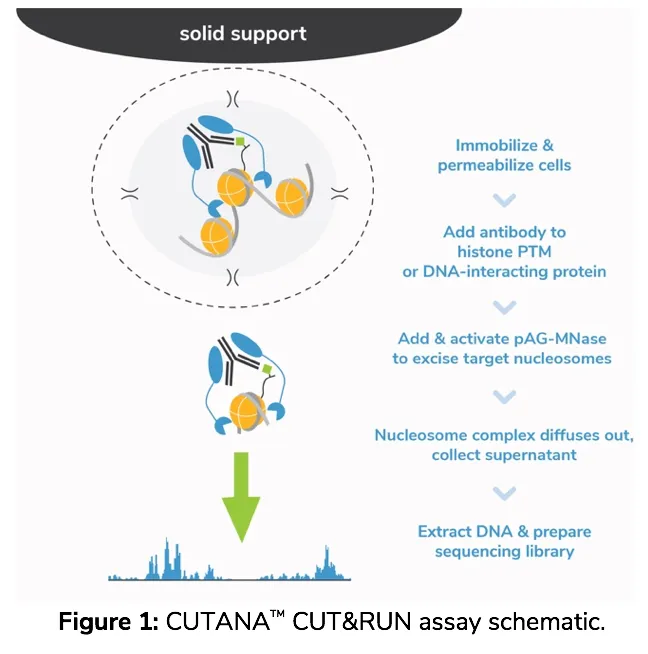
For those new to CUT&RUN, check out our recent blog post detailing CUT&RUN methodology as well as our video summary of the major advantages of CUT&RUN vs. ChIP-seq.
Are Transcription Factors Compatible with CUT&RUN?
CUT&RUN has been applied to countless histone post-translational modifications (PTMs) with great success. Additionally, published reports have shown that CUT&RUN can be used to derive high-resolution genomic profiles of chromatin associated proteins, including transcription factors (TFs) such as CTCF 2, 4, 5, FoxA1 / FoxA2 5, OCT4 4, 5, and others 6-8. However, anecdotal reports have emerged that suggest current CUT&RUN protocols do not consistently work for TFs. Indeed, we have heard this feedback from many collaborators and customers, including some interactions on Twitter!
TFs are a major focus of epigenetics research, and the ability to use CUT&RUN to map TFs would have a significant impact on the chromatin field. Recognizing this need, EpiCypher took on the task of developing an upgraded CUT&RUN protocol, optimized for robust profiling of TFs, histone PTMs, and other chromatin-interacting proteins.
A New and Improved CUT&RUN Protocol

In the new protocol, highly reproducible results are enabled by adoption of a user-friendly 8-well strip-tube format, which simplifies sample handling and supports multichannel pipetting or assay automation. An added benefit of this updated protocol is that it allows increased experimental throughput, taking advantage of one of the major benefits of CUT&RUN over ChIP-seq: reduced sequencing depth requirements (3-5 million reads per sample compared to ~30 million in ChIP-seq). Thus, with this newfound technology, scientists will be able to run more experiments, more samples and additional replicates at a lower per-reaction cost, enabling a more comprehensive analysis of complex biological mechanisms that have been inaccessible by existing approaches.
Of note, we offer a magnetic stand (Catalog No. #10-0008) that can hold two 8 strip-tubes, and is identical to the magnet we used for assay optimization! In addition, we are also selling ConA magnetic beads (Catalog No. #21-1401), and an extensively validated H3K4me3 positive control antibody (Catalog No. 13-0041) which are directly compatible with our new CUT&RUN protocol.
Before you dive into our protocol, we want to highlight a few of the Frequently Asked Questions (FAQs) that might be helpful in setting up your CUT&RUN experiment.
What is the best way to know if a CUT&RUN experiment worked prior to sequencing?
Results from challenging cell inputs / targets may be ambiguous, so EpiCypher recommends including positive (e.g. H3K4me3 EpiCypher Catalog No. 13-0041) and negative (IgG or no antibody) controls in every experiment. If the QC checks for the positive control perform as expected, then you are all set for library prep and sequencing! If sequencing results for challenging cell inputs / targets are not satisfactory further optimization may be necessary (e.g. cell type and / or number, digitonin permeabilization, antibody concentration / alternate vendors, etc).
Can residual E. coli in the pAG-MNase prep be used for sample input normalization? What spike-in DNA control does EpiCypher recommend?
Carry-over E. coli DNA is present in EpiCypher’s pAG-MNase preps. However, at a typical sequencing depth of 3-5 million reads, too few E. coli DNA fragments (~hundreds) are recovered for reliably computing sample normalization. Thus, we recommend considering other normalization strategies, such as exogenous DNA spike-in (E. coli / yeast / fly) or recombinant nucleosome spike-ins (in development!).
What types of cell inputs are compatible with CUT&RUN?
EpiCypher has tested and validated our CUT&RUN protocol using whole cells and nuclei, derived from mammalian suspension and adherent cancer lines. We have not yet directly tested primary tissue, FACS sorted, or immune cells*. However, a number of groups have successfully performed CUT&RUN on human and mouse primary tissue 6, 9-11, FACS sorted 4 and immune cells 12, 13.
*Note that lectins (e.g. ConA-beads) play a role in the innate immune system and immune cells types may be inadvertently stimulated via binding to ConA-beads. To circumvent this potential problem, EpiCypher recommends using nuclei 4 or a crosslinking strategy 14 in CUT&RUN.
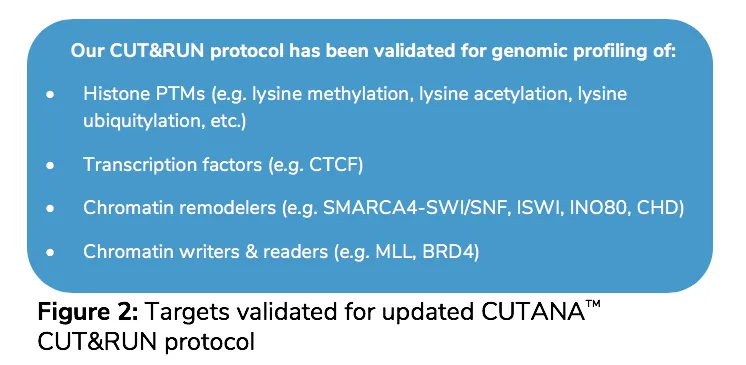
Does EpiCypher’s CUT&RUN protocol work on non-PTM targets?
Yes! Our updated protocol has been used to generate non-PTM CUT&RUN data, including CTCF, BRD4, and SMARCA4 (BRG1). No protocol modifications were necessary to generate these data since the columns we use to purify the CUT&RUN enriched chromatin fragments retains >50bp DNA.
References
1. Meers MP, et al. Improved CUT&RUN chromatin profiling tools. Elife, 2019. 8: p. (PubMed PMID: 31232687) (PMC6598765)
2. Skene PJ, et al. Targeted in situ genome-wide profiling with high efficiency for low cell numbers. Nat Protoc, 2018. 13(5): p. 1006-19. (PubMed PMID: 29651053)
3. Skene PJ, Henikoff S. An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. Elife, 2017. 6: p. (PubMed PMID: 28079019) (PMC5310842)
4. Hainer SJ, et al. Profiling of Pluripotency Factors in Single Cells and Early Embryos. Cell, 2019. 177(5): p. 1319-29 e11. (PubMed PMID: 30955888) (PMC6525046)
5. Meers MP, et al. Pioneer Factor-Nucleosome Binding Events during Differentiation Are Motif Encoded. Mol Cell, 2019. 75(3): p. 562-75 e5. (PubMed PMID: 31253573) (PMC6697550)
6. Liu N, et al. Direct Promoter Repression by BCL11A Controls the Fetal to Adult Hemoglobin Switch. Cell, 2018. 173(2): p. 430-42 e17. (PubMed PMID: 29606353) (PMC5889339)
7. Rothstein M, Simoes-Costa M. Heterodimerization of TFAP2 pioneer factors drives epigenomic remodeling during neural crest specification. Genome Res, 2020. 30(1): p. 35-48. (PubMed PMID: 31848212)
8. Zhu Q, et al. CUT&RUNTools: a flexible pipeline for CUT&RUN processing and footprint analysis. Genome Biol, 2019. 20(1): p. 192. (PubMed PMID: 31500663) (PMC6734249)
9. de Bock CE, et al. HOXA9 Cooperates with Activated JAK/STAT Signaling to Drive Leukemia Development. Cancer Discov, 2018. 8(5): p. 616-31. (PubMed PMID: 29496663)
10. Janssens DH, et al. Automated in situ chromatin profiling efficiently resolves cell types and gene regulatory programs. Epigenetics & chromatin, 2018. 11(1): p. 74. (PubMed PMID: 30577869) (PMC6302505)
11. Uyehara CM, McKay DJ. Direct and widespread role for the nuclear receptor EcR in mediating the response to ecdysone in Drosophila. Proc Natl Acad Sci U S A, 2019. 116(20): p. 9893-902. (PubMed PMID: 31019084) (PMC6525475)
12. Mathsyaraja H, et al. Max deletion destabilizes MYC protein and abrogates Emicro-Myc lymphomagenesis. Genes & development, 2019. p. (PubMed PMID: 31395740)
13. Roth TL, et al. Reprogramming human T cell function and specificity with non-viral genome targeting. Nature, 2018. 559(7714): p. 405-9. (PubMed PMID: 29995861) (PMC6239417)
14. Zheng XY, Gehring M. Low-input chromatin profiling in Arabidopsis endosperm using CUT&RUN. Plant Reprod, 2019. 32(1): p. 63-75. (PubMed PMID: 30719569)
