CUT&RUN Just Got Easier
- Ellen Weinzapfel

EpiCypher is leading the development of user-friendly solutions for emerging epigenomic mapping technologies. Our recently launched CUTANA™ CUT&RUN Library Prep Kits, powered by NEBNext® reagents from New England Biolabs® (NEB®), are the first library prep kits specifically optimized for CUT&RUN assays. These library prep kits seamlessly integrate with our CUTANA™ CUT&RUN Kit and Protocol, providing a streamlined workflow from cells to sequence-ready libraries.
Why did we make CUTANA Library Prep Kits, and what makes them so unique? Read on to learn more!
The current complexities of CUT&RUN library prep
CUT&RUN (Cleavage Under Targets and Release Using Nuclease) is a powerful new approach to map chromatin features with dramatically reduced cell input and sequencing requirements compared to ChIP-seq (learn more). But CUT&RUN is only the first step of the experiment! Following CUT&RUN, researchers must prepare their enriched target DNA for next generation sequencing (NGS) using a library prep kit and reagents.
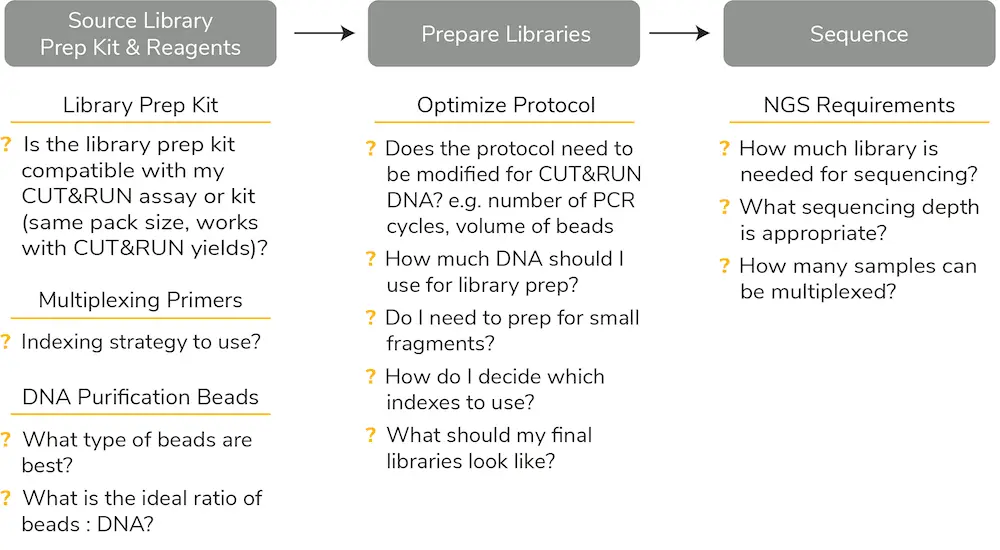
NGS library prep is a multi-step method that requires navigating a complex web of questions before starting the experiment (Figure 1). Users must determine which library prep kit to purchase, the indexing strategy and multiplexing primers they want to use, and the type / ratio of DNA purification beads they need.
It is also important to optimize the library prep protocol for the upstream assay, as different mapping assays generate different input amounts. Because CUT&RUN is an ultra-sensitive assay with high signal-to-noise, low background, and reduced cell inputs, it intrinsically generates low DNA yields compared to ChIP. All these factors – and more – need to be considered when planning CUT&RUN library prep (Figure 1).
Why are there so many decisions? To maximize the application of their products, vendors offer general “DNA library prep kits” that can be combined with various multiplexing strategies and adapted for assay-specific inputs. These kits aren’t unique to CUT&RUN assays – in fact, few companies offer assay-specific library prep kits.
Despite the versatility of this product strategy, being “everything for everyone” has several drawbacks:
- The number of options makes it confusing to identify the best library prep method for your experiment.
- The generic nature of library prep protocols complicates integration with new profiling strategies, such as CUT&RUN.
- You have to spend extra money to source all the tools you need, since assay kits, library prep kits, DNA purification beads, and associated reagents come in different pack sizes from multiple vendors.
Can’t we just use ChIP-seq library prep protocols? CUT&RUN users have relied on ChIP-seq library prep kits or generic DNA library prep kits adapted to lower DNA input. But modifying ChIP-seq library prep for ultra-sensitive CUT&RUN assays has been difficult for many reasons (see Figure 1):
- Lower yields from CUT&RUN challenges the use of ChIP-seq library prep protocols. Are lower inputs within the sensitivity limits of the library prep kit? Should the amount of library prep enzymes, sequencing adapter, or indexing primer be reduced? Are additional PCR cycles needed to ensure there is sufficient library for NGS?
- Unclear how to confirm high quality of CUT&RUN libraries. Fragment distribution analysis (e.g. by Agilent Bioanalyzer®) is an important quality control step during library prep, and data are used to pool samples for NGS. Libraries using CUT&RUN DNA appear different compared to ChIP-seq libraries, complicating this step.
- Lack of guidance on NGS requirements for CUT&RUN. One of the core advantages of CUT&RUN is high quality data at low sequencing depth. But many users continue to use 20 million (or more) reads per reaction, limiting multiplexing capabilities and increasing NGS costs.
The combination of decisions and assay optimization required for CUT&RUN library prep is intimidating, even for seasoned researchers, and has slowed the adoption of CUT&RUN by some groups.
EpiCypher decided to develop CUT&RUN assay-specific Library Prep Kits to provide a comprehensive solution to the concerns outlined above. The resulting CUTANA workflow makes it easier than ever to perform CUT&RUN assays, with major gains in throughput and reduced costs.
Now that we have covered “why” we made our Library Prep Kits, let’s dive into “how” we did it.
CUTANA™ CUT&RUN Library Prep Kits
Our strategy for developing the CUTANA CUT&RUN Library Prep Kit was straightforward: find the best library prep reagents, optimize their application for CUT&RUN DNA inputs, and then package all key reagents and the optimized protocol together in one box. The primary goal was to simplify the workflow for CUTANA users by providing a reasonably priced, all-inclusive library prep kit expressly validated for CUT&RUN.
These rigorous efforts led to our CUTANA CUT&RUN Library Prep Kits, powered by NEB’s NEBNext reagents and multiplexing oligos. NEB has been developing reagents for library preparation for over 12 years, and in our hands, these reagents generate the highest quality CUT&RUN data. In fact, during kit development, we tested all the top library kits on the market and found that NEBNext generated the most reliable data with highly sensitivity – a perfect match for our CUTANA CUT&RUN assays.
We subsequently partnered with NEB to optimize NEBNext reagents for our CUT&RUN assays. The resulting library prep kits & protocol:
- Include all the reagents needed for CUT&RUN library prep – including NEBNext reagents, NEB multiplexing oligos, and DNA purification beads. This provides massive advantages compared to competitor kits, while still offering significant cost savings (Figure 2).
- Have been uniquely optimized for CUT&RUN enriched DNA. Our Library Prep Kit Manual highlights key protocol steps, offers guidance on index primer selection, and has an extensive FAQ section.
- Generate robust CUT&RUN sequencing data for a range of input amounts and targets. Our kits have been validated for histone post-translational modifications (e.g. H3K4me3, H3K27me3) and chromatin associated proteins (e.g. CTCF) using as few as 0.5 ng CUT&RUN DNA as input.
- Enable multiplexing of up to 96 CUT&RUN libraries in a single NGS run. The primer indexing strategy allows each 48-reaction kit to be multiplexed in a single NGS run. The two versions of the library prep kit (14-1001 & 14-1002) contain distinct sets of indexing primers, allowing up to 96 CUT&RUN libraries to be multiplexed when kits are used together.

Two CUTANA™ Kits: An end-to-end solution for CUT&RUN chromatin mapping
The most important advantage of the CUTANA Library Prep Kits are their seamless integration with the CUTANA CUT&RUN Kit and Do-It-Yourself Protocol, thus providing an end-to-end solution for all users (Figure 3). Benefits of combining the CUT&RUN Kit and Library Prep Kits include:
- Optimized kits contain all reagents to go from cells to NGS libraries.
- All kits are designed for 48 reactions, simplifying workflow set up and minimizing reagent waste.
- Each protocol is compatible with multi-channel pipettors for improved reliability and increased throughput.
- Both kits come with a benchtop-friendly Quick-Start Card and a detailed manual including troubleshooting tips, FAQs, and much more!
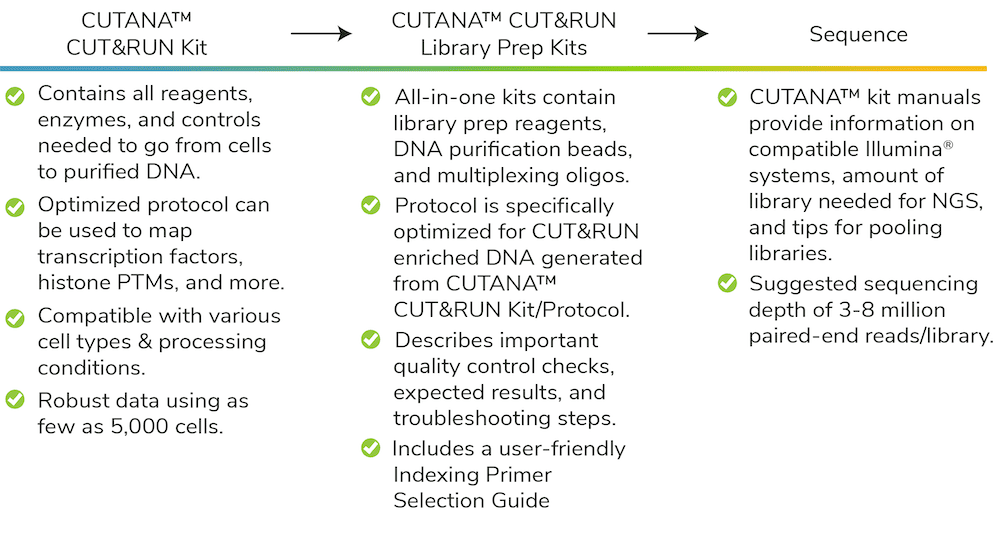
Each reagent in these kits has been meticulously tested and verified by EpiCypher scientists, and our protocols are robust for various targets and inputs. The resulting CUTANA workflow (Figure 3) makes CUT&RUN accessible to users of all experience levels, which will improve the quality of chromatin mapping and transform epigenomics research.
Our CUTANA CUT&RUN Kit and Library Kits are available now, along with a collection of rigorously validated antibodies, controls, and other supporting reagents. Shop here or inquire using the form below!
