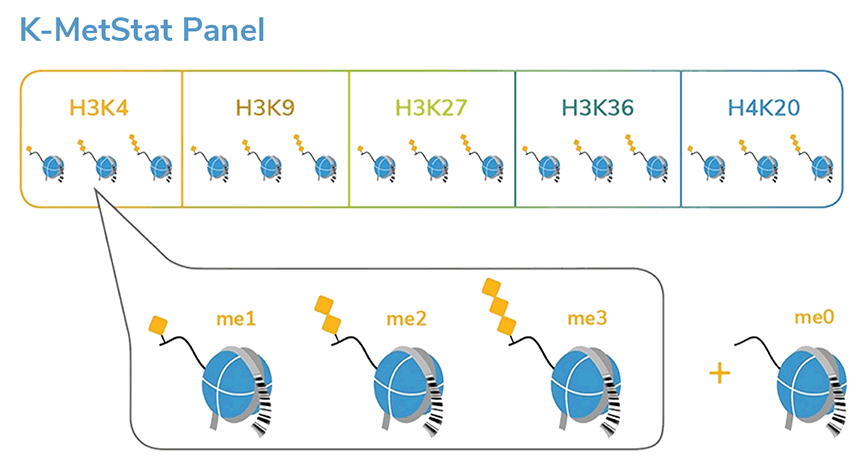
SNAP-ChIP® K-MetStat Panel
$365.00
SKU: 19-1001
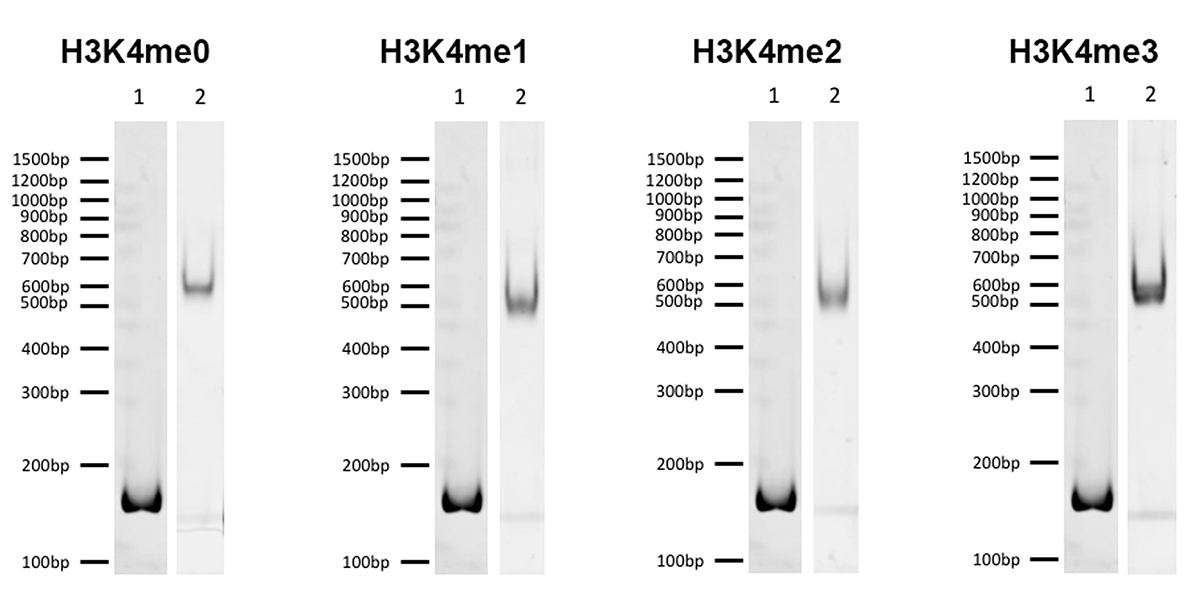
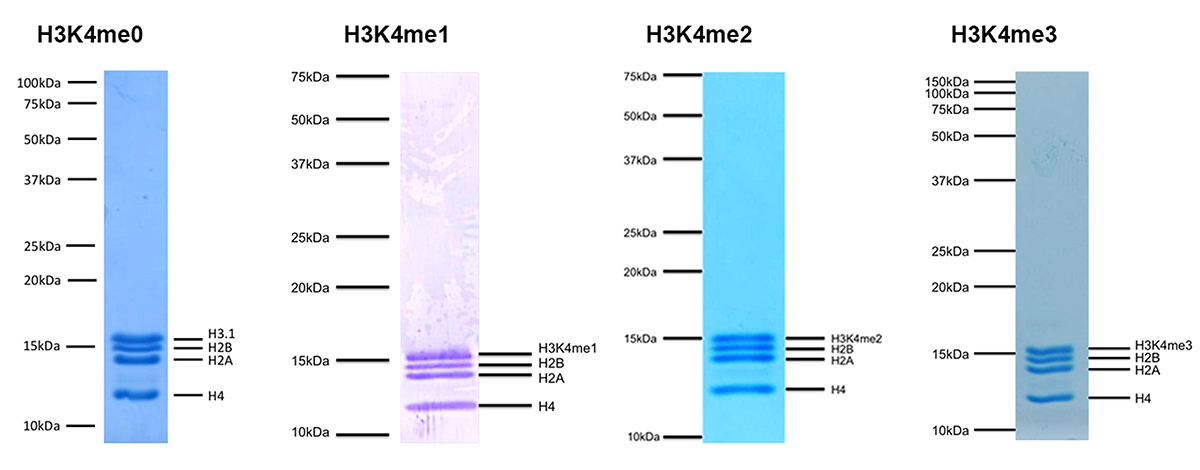
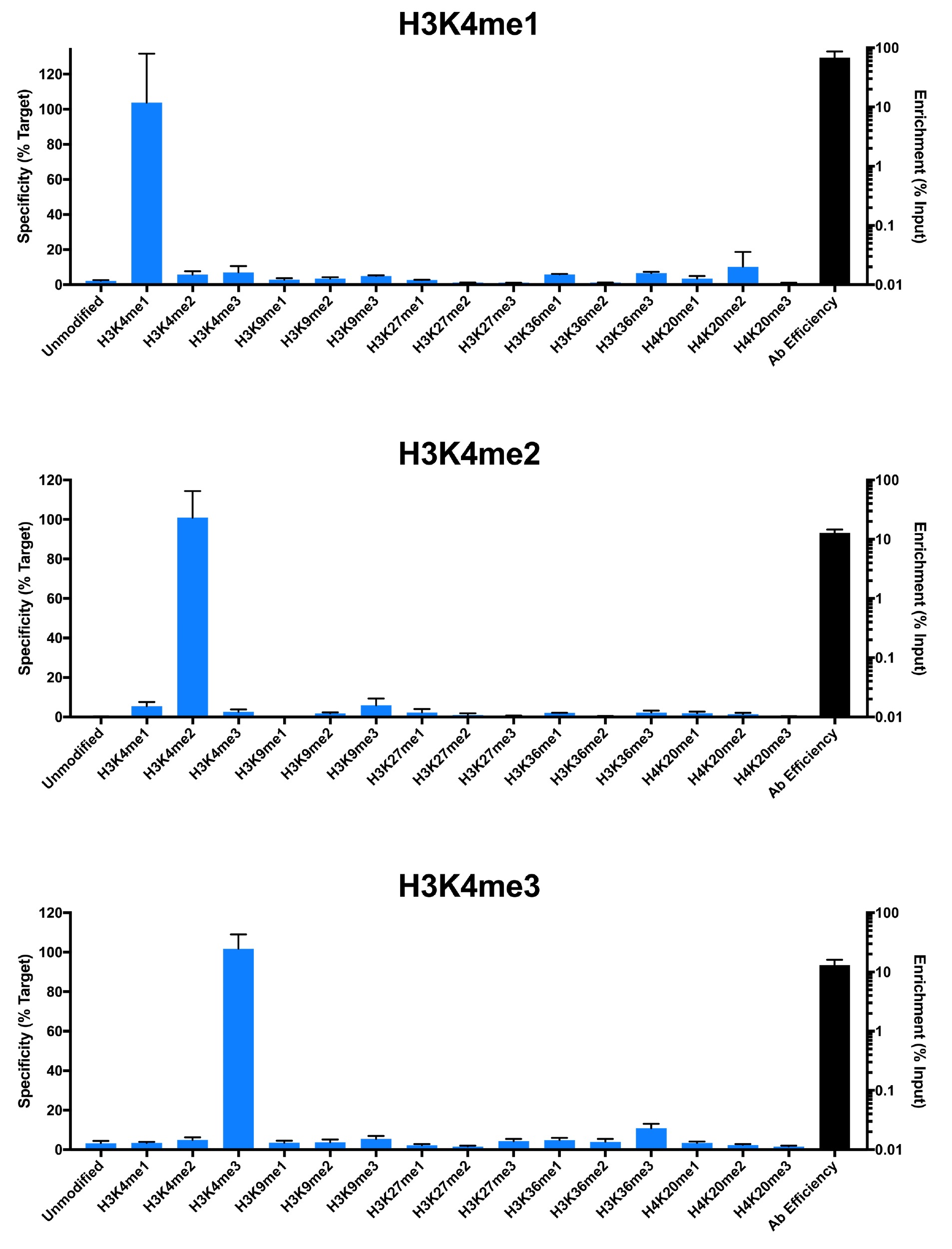
{"sku":"19-1001","url":"https://www.epicypher.com/products/nucleosomes/snap-chip-k-metstat-panel","add_this":[{"service":"facebook","annotation":""},{"service":"email","annotation":""},{"service":"print","annotation":""},{"service":"twitter","annotation":""},{"service":"linkedin","annotation":""}],"gtin":"","id":"411","bulk_discount_rates":[],"can_purchase":true,"meta_description":"SNAP-ChIP K-MetStat Panel Spike-in Normalization Control for Chromatin IP (ChIP) and ChIP-Seq","category":["Nucleosomes","Nucleosomes/SNAP-ChIP<sup>®</sup> Spike-ins"],"main_image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/411/282/SNAP_Chip__53961.1569012494.png?c=2","alt":"SNAP-ChIP® K-MetStat Panel"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=411","shipping":{"calculated":true},"num_reviews":0,"weight":"0.01 LBS","description":"<table border=\"0\" cellpadding=\"2\" cellspacing=\"2\" width=\"100%\">\n\t<tbody>\n\t\t<tr valign=\"top\">\n\t\t\t<td>\n\t\t\t<table border=\"0\" cellpadding=\"1\" cellspacing=\"1\" height=\"1490\" width=\"90%\">\n\t\t\t\t<tbody>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td valign=\"top\">\n\t\t\t\t\t\t<p></p>\n\n\t\t\t\t\t\t\n\t\t\t\t\t\t<span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><b><b>SNAP-ChIP K-MetStat Panel</b> Description:</b> A panel of distinctly modified mononucleosomes assembled from recombinant human histones expressed in <em>E. coli</em> (two each of histones H2A, H2B, H3 and H4; accession numbers: H2A-P04908; H2B-O60814; H3.1-P68431 or H3.2-Q71DI3*; H4-P62805) wrapped by 147 base pairs of barcoded Widom 601 positioning sequence DNA. The panel is comprised of a pool of 1 unmodified plus 15 histone H3 or H4 post-translational modifications (PTMs, created by a proprietary semi-synthetic method): H3K4me1, H3K4me2, H3K4me3, H3K9me1, H3K9me2, H3K9me3, H3K27me1, H3K27me2, H3K27me3, H3K36me1, H3K36me2, H3K36me3, H4K20me1, H4K20me2 and H4K20me3. Each distinctly modified nucleosome is distinguishable by a unique sequence of DNA ("barcode") at the 3' end that can be deciphered by qPCR or next-generation sequencing. Each of the 16 nucleosomes in the pool is wrapped by 2 distinct DNA species, each containing a distinct barcode ("A" and "B", see SNAP-ChIP Manual) allowing for an internal technical replicate.</span></span>\n\n\t\t\t\t\t\t<p><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><span style=\"color: orange !important;\"><b>Learn more about SNAP-ChIP Certified Antibodies <a href=\"https://www.epicypher.com/snap-chip-certified-antibodies/\">here</a></b></span></span></span></p>\n\n\t\t\t\t\t\t<p><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\">* Histone H3.2 contains a Cys to Ala mutation at position 110. </span></span><br />\n\t\t\t\t\t\t</p>\n\t\t\t\t\t\t</td>\n\t\t\t\t\t</tr>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td valign=\"top\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><b><b>SNAP-ChIP K-MetStat Panel</b> Formulation:</b> Purified recombinant mononucleosomes, containing a mixture of 16 (1 unmodified plus 15 unique) H3 and H4 PTMs in 10 mM sodium cacodylate, pH 7.5, 100 mM NaCl, 1 mM EDTA, 50% glycerol (w/v), 1x Protease Inhibitor cocktail, 100 ug/mL BSA, 10 mM β-mercaptoethanol. Molarity = 0.6 nM. MW = ~199382.1 Da (average MW of all 16 nucleosomes).</span></span><br /><br />\n\t\t\t\t\t\t</td>\n\t\t\t\t\t</tr>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td valign=\"top\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><b>SNAP-ChIP K-MetStat Panel Storage and Stability:</b> Stable for six months at -20°C from date of receipt.</span></span><br /><br />\n\t\t\t\t\t\t</td>\n\t\t\t\t\t</tr>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td valign=\"top\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><b>SNAP-ChIP K-MetStat Panel Application Notes:</b> SNAP-ChIP K-MetStats are highly purified recombinant mononucleosomes and are suitable for use as spike-in controls for ChIP reactions, for antibody specificity testing or for effector protein binding experiments. SNAP-ChIP K-MetStats from EpiCypher do not contain detectable levels of free DNA which could alter assayed activities. See manual for more information. SNAP-ChIP can easily be introduced into existing ChIP and ChIP-seq workflows for experiment normalization and monitoring of technical variability. </span></span>\n\t\t\t\t\t\t<p><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><span style=\"color: orange !important;\"><b>Request a quote <a href=\"https://www.epicypher.com/request-a-quote/\">here</a></b></span></span></span></p>\n\t\t\t\t\t\t</td>\n\t\t\t\t\t</tr>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td valign=\"top\">\n\t\t\t\t\t\t<p><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><b>Background References</b>:</span></span><br>\n\t\t\t\t\t\t <span style=\"font-family:arial,helvetica,sans-serif;\">Grzybowski A et al. 2015 Mol Cell 58: 886 - 899 <a href=\"https://www.ncbi.nlm.nih.gov/pubmed/26004229\" title=\"Calibrating ChIP-Seq with Nucleosomal Internal Standards to Measure Histone Modification Density Genome Wide\" target=\"_new\">LINK</a></span>\t\t\t\t\t\t</p>\n\t\t\t\t\t\t<p><span style=\"font-family:arial,helvetica,sans-serif;\"><span style=\"font-size:12pt;\"><b>Product References</b>:<br>\n\t\t\t\t\t\t</span></span><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\">Shah RN et al. 2018 Mol. Cell 72: 162–177 <a href=\"https://www.ncbi.nlm.nih.gov/pubmed/30244833\" title=\"Examining the Roles of H3K4 Methylation States with Systematically Characterized Antibodies\" target=\"_new\">LINK</a></span></span>\t\t\t\t\t\t<br>\n <span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\">Janssen A et al. 2018 Genes Dev. 33: 103-115 <a href=\"https://pubmed.ncbi.nlm.nih.gov/30578303/\">LINK </a><br>\n Yeganeh M et al. 2018 Nucl. Acids Res. <a href=\"https://pubmed.ncbi.nlm.nih.gov/30597109/\" title=\"Differential regulation of RNA polymerase III genes during liver regeneration\" target=\"_new\">LINK</a></span></span>\t\t\t\t\t\t<br />\n\t\t\t\t\t\t<br />\n\t\t\t\t\t </p></td>\n\t\t\t\t\t</tr>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td valign=\"top\">\n\t\t\t\t\t\t<p><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><b>SNAP-ChIP K-MetStat Panel Additional Resources and Files:</b></span></span></p>\n\n\t\t\t\t\t\t<table border=\"0\" cellpadding=\"0\" cellspacing=\"10\" height=\"447\" width=\"450\">\n\t\t\t\t\t\t\t<tbody>\n\t\t\t\t\t\t\t\t<tr>\n\t\t\t\t\t\t\t\t\t<td style=\"height:60px\" valign=\"top\" width=\"350\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Technical datasheet </strong></span></span></td>\n\t\t\t\t\t\t\t\t\t<td align=\"center\" valign=\"top\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/documents/tds/19-1001.pdf\" target=\"_new\"><img alt=\"19-1001 Datasheet\" height=\"40\" src=\"/content/documents/tds/icon.png\" width=\"30\" /></a></span></span></td>\n\t\t\t\t\t\t\t\t</tr>\n\t\t\t\t\t\t\t\t<tr>\n\t\t\t\t\t\t\t\t\t<td style=\"height:60px\" valign=\"top\" width=\"350\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>SNAP-ChIP K-MetStat Manual</strong></span></span></td>\n\t\t\t\t\t\t\t\t\t<td align=\"center\" valign=\"top\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/documents/snapchip/SNAP-ChIP_manual.pdf\" target=\"_new\"><img alt=\"SNAP-ChIP K-MetStat manual.pdf\" height=\"40\" src=\"/content/documents/tds/icon.png\" width=\"30\" /></a></span></span></td>\n\t\t\t\t\t\t\t\t</tr>\n\t\t\t\t\t\t\t\t<tr>\n\t\t\t\t\t\t\t\t\t<td style=\"height:60px\" valign=\"top\" width=\"350\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>SNAP-ChIP Brochure</strong></span></span></td>\n\t\t\t\t\t\t\t\t\t<td align=\"center\" valign=\"top\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/docs/SNAP-ChIP.pdf\" target=\"_new\"><img alt=\"SNAP-ChIP K-MetStat manual.pdf\" height=\"40\" src=\"/content/documents/tds/icon.png\" width=\"30\" /></a></span></span></td>\n\t\t\t\t\t\t\t\t</tr>\n\t\t\t\t\t\t\t\t<tr>\n\t\t\t\t\t\t\t\t\t<td style=\"height:60px\" valign=\"top\" width=\"350\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>SNAP-ChIP K-MetStat Full Panel Information: </strong>Forward/reverse barcodes, nucleosomal DNA sequences, qPCR primer sequences, TaqMan probe sequence</span></span></td>\n\t\t\t\t\t\t\t\t\t<td align=\"center\" valign=\"top\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/documents/snapchip/ SNAP-ChIP_K-MetStat_19-1001_Barcode-Sequence-Primer_Info-SHORT_ver1.1.xlsx\" target=\"_new\"><img alt=\"19-1001 Datasheet\" height=\"40\" src=\"/content/documents/excelicon.png\" width=\"30\" /></a></span></span></td>\n\t\t\t\t\t\t\t\t</tr>\n\t\t\t\t\t\t\t\t<tr>\n\t\t\t\t\t\t\t\t\t<td style=\"height:60px\" valign=\"top\" width=\"350\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>SNAP-ChIP K-MetStat qPCR Primer Sequences</strong> - Formatted for easy upload for ordering from common vendors (<em>e.g.</em> IDT)</span></span></td>\n\t\t\t\t\t\t\t\t\t<td align=\"center\" valign=\"top\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/documents/snapchip/SNAP-ChIP_K-MetStat_qPCR_Primers_BarcodeA.xlsx\" target=\"_new\"><img alt=\"19-1001 Datasheet\" height=\"40\" src=\"/content/documents/excelicon.png\" width=\"30\" /></a></span></span></td>\n\t\t\t\t\t\t\t\t</tr>\n\t\t\t\t\t\t\t\t<tr>\n\t\t\t\t\t\t\t\t\t<td style=\"height:60px\" valign=\"top\" width=\"350\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Fasta formatted file of K-MetStat barcode Sequences</strong> - For alignment of NGS sequencing data</span></span></td>\n\t\t\t\t\t\t\t\t\t<td align=\"center\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/documents/snapchip/KmetStat_barcoded_fasta.txt.rtf\" target=\"_new\"><img alt=\"19-1001 fasta\" height=\"40\" src=\"/content/documents/excelicon.png\" width=\"30\" /></a></span></td>\n\t\t\t\t\t\t\t\t</tr>\n\t\t\t\t\t\t\t</tbody>\n\t\t\t\t\t\t</table>\n\t\t\t\t\t\t</td>\n\t\t\t\t\t</tr>\n\t\t\t\t</tbody>\n\t\t\t</table>\n\t\t\t</td>\n\t\t\t<td>\n\t\t\t<table border=\"0\" cellpadding=\"3\" cellspacing=\"3\" width=\"100%\">\n\t\t\t\t<tbody>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td align=\"center\"><span style=\"font-size:11pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/images/nucleosome/K-MetStat_Graphic.gif\" target=\"_new\"><img alt=\"K-MetStat Panel\" src=\"/content/images/nucleosome/K-MetStat_Graphic.gif\" width=\"250\" /></a></span></span></td>\n\t\t\t\t\t</tr>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td align=\"left\" valign=\"top\"><span style=\"font-size:11pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>SNAP-ChIP K-MetStat Panel:</strong> Schematic representation of nucleosomes comprising the K-MetStat Panel: H3K4me1, H3K4me2, H3K4me3, H3K9me1, H3K9me2, H3K9me3, H3K27me1, H3K27me2, H3K27me3, H3K36me1, H3K36me2, H3K36me3, H4K20me1, H4K20me2 and H4K20me3 and me0 (unmodified).<br />\n\t\t\t\t\t\t<strong>(Click image to enlarge) </strong></span></span>\n\t\t\t\t\t\t<p></p>\n\t\t\t\t\t\t</td>\n\t\t\t\t\t</tr>\n\t\t\t\t</tbody>\n\t\t\t</table>\n\n\t\t\t<table border=\"0\" cellpadding=\"3\" cellspacing=\"3\" width=\"100%\">\n\t\t\t\t<tbody>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td align=\"center\"><span style=\"font-size:11pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/images/products/nucleosomes/19-1001-DNA-gel.jpg\" target=\"_new\"><img alt=\"19-1001 DNA Gel Data\" src=\"/content/images/products/nucleosomes/19-1001-DNA-gel.jpg\" width=\"250\" /></a></span></span></td>\n\t\t\t\t\t</tr>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td align=\"left\" valign=\"top\"><span style=\"font-size:11pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>DNA Gel Data:</strong> Representative images for SNAP-ChIP K-MetStats (H3K4me0 = unmodified, H3K4me2, H3K4me3) run on a native PAGE gel and stained with ethidium bromide to visualize DNA. Lane 1: Free 147bp DNA used in nuclesome assembly (100 ng). Lane 2: Intact nucleosomes (200 ng) showing lack of free DNA. Identical experiments were performed for the entire K-MetStat Panel.<br />\n\t\t\t\t\t\t<strong>(Click image to enlarge) </strong></span></span>\n\t\t\t\t\t\t<p></p>\n\t\t\t\t\t\t</td>\n\t\t\t\t\t</tr>\n\t\t\t\t</tbody>\n\t\t\t</table>\n\n\t\t\t<table border=\"0\" cellpadding=\"3\" cellspacing=\"3\" width=\"100%\">\n\t\t\t\t<tbody>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td align=\"center\"><span style=\"font-size:11pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/images/products/nucleosomes/19-1001-protein-gel.jpg\" target=\"_new\"><img alt=\"19-1001 Protein Gel Data\" src=\"/content/images/products/nucleosomes/19-1001-protein-gel.jpg\" width=\"250\" /></a></span></span></td>\n\t\t\t\t\t</tr>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td align=\"left\" valign=\"top\"><span style=\"font-size:11pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Protein Gel Data:</strong> Representative Coomassie stained PAGE gel for SNAP-ChIP K-MetStats (2 µg each of unmodified, H3K4me1, H3K4me2, H3K4me3) to demonstrate the purity of the histones in the preparation. Sizes of molecular weight markers and positions of the core histones (H2A, H2B, H3 and H4) are indicated. Identical experiments were performed for the remainder of the K-MetStat Panel.<br />\n\t\t\t\t\t\t<strong>(Click image to enlarge) </strong></span></span>\n\t\t\t\t\t\t<p></p>\n\t\t\t\t\t\t</td>\n\t\t\t\t\t</tr>\n\t\t\t\t</tbody>\n\t\t\t</table>\n\n\t\t\t<table border=\"0\" cellpadding=\"3\" cellspacing=\"3\" width=\"100%\">\n\t\t\t\t<tbody>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td align=\"center\"><span style=\"font-size:11pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/images/products/nucleosomes/19-1001-ChIP-data.jpg\" target=\"_new\"><img alt=\"19-1001 ChIP Data\" src=\"/content/images/products/nucleosomes/19-1001-ChIP-data.jpg\" width=\"250\" /></a></span></span></td>\n\t\t\t\t\t</tr>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td align=\"left\" valign=\"top\">\n\t\t\t\t\t\t<p><span style=\"font-size:11pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>ChIP Data:</strong> Representative images for SNAP-ChIP K-MetStats (unmodified, H3K4me1, H3K4me2, H3K4me3) assayed in a chromatin immunoprecipitation (ChIP) experiment using commercially available ChIP grade antibodies (3 µg, n = 3). Quantitative Real-Time PCR (qPCR) for the DNA barcodes corresponding to unmodified (H3K4me0), H3K4me1, H3K4me2, and H3K4me3 nucleosomes show recovery of the barcodes corresponding to the expected antibody target. Identical experiments were performed for the remainder of the K-MetStat Panel (H3K9me1, H3K9me2, H3K9me3, H3K27me1, H3K27me2, H3K27me3, H3K36me1, H3K36me2, H3K36me3, H4K20me1, H4K20me2 and H4K20me3).</span></span></p>\n\n\t\t\t\t\t\t<p><span style=\"font-size:11pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>N.B.</strong>-SNAP-ChIP is extremely useful for determining antibody cross-reactivity with off-target modifications. The cross-reactivity observed above (K4me1 & K4me2) is not indicative of a failure in the SNAP-ChIP procedure, but in the antibody to specifically recognize its target.<br />\n\t\t\t\t\t\t<strong>(Click image to enlarge) </strong></span></span></p>\n\n\t\t\t\t\t\t<p></p>\n\t\t\t\t\t\t</td>\n\t\t\t\t\t</tr>\n\t\t\t\t</tbody>\n\t\t\t</table>\n\n\t\t\t<table border=\"0\" cellpadding=\"3\" cellspacing=\"3\" width=\"100%\">\n\t\t\t\t<tbody>\n\t\t\t\t</tbody>\n\t\t\t</table>\n\t\t\t</td>\n\t\t</tr>\n\t</tbody>\n</table>\n\n","tags":[],"warranty":"","price":{"without_tax":{"formatted":"$365.00","value":365,"currency":"USD"},"tax_label":"Sales Tax"},"detail_messages":"","availability":"","page_title":"SNAP-ChIP K-MetStat Panel Spike-in Normalization Control for Chromatin IP (ChIP) and ChIP-Seq","cart_url":"https://www.epicypher.com/cart.php","max_purchase_quantity":0,"mpn":"","upc":null,"options":[{"id":682,"type":"Configurable_PickList_Set","display_name":"Select Pack Size:","required":true,"condition":true,"state":"variant_option","values":[{"label":"20 µl","id":111,"data":"20 µl","selected":true},{"label":"200 µl","id":106,"data":"200 µl","selected":false}],"partial":"set-radio"}],"related_products":[{"id":555,"sku":null,"name":"SNAP-ChIP® OncoStat Panel","url":"https://www.epicypher.com/products/nucleosomes/snap-chip-oncostat-panel","availability":"","rating":null,"brand":{"name":null},"category":["Nucleosomes","Nucleosomes/SNAP-ChIP<sup>®</sup> Spike-ins"],"summary":" \n\n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/555/399/OncoStat__15594.1569012525.png?c=2","alt":"SNAP-ChIP® OncoStat Panel"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/555/399/OncoStat__15594.1569012525.png?c=2","alt":"SNAP-ChIP® OncoStat Panel"}],"date_added":"31st Jul 2018","pre_order":false,"show_cart_action":true,"has_options":true,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":null,"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"price":{"without_tax":{"currency":"USD","formatted":"$365.00","value":365},"price_range":{"min":{"without_tax":{"currency":"USD","formatted":"$365.00","value":365},"tax_label":"Sales Tax"},"max":{"without_tax":{"currency":"USD","formatted":"$2,795.00","value":2795},"tax_label":"Sales Tax"}},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=555"},{"id":626,"sku":"13-0028","name":"Histone H3K4me3 Antibody, SNAP-ChIP Certified *DISCONTINUED*","url":"https://www.epicypher.com/products/antibodies/snap-chip-certified-antibodies/histone-h3k4me3-antibody-snap-chip-certified","availability":"","rating":null,"brand":{"name":null},"category":[],"summary":"\n \n \n \n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/626/578/Screen_Shot_2018-09-25_at_2.00.27_PM__52796.1538135192.png?c=2","alt":"Histone H3K4me3 Antibody, SNAP-ChIP Certified *DISCONTINUED*"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/626/578/Screen_Shot_2018-09-25_at_2.00.27_PM__52796.1538135192.png?c=2","alt":"Histone H3K4me3 Antibody, SNAP-ChIP Certified *DISCONTINUED*"}],"date_added":"24th Sep 2018","pre_order":false,"show_cart_action":false,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":452,"name":"Pack Size","value":"100 μg"}],"num_reviews":null,"weight":{"formatted":"0.00 LBS","value":0},"demo":false,"add_to_wishlist_url":"/wishlist.php?action=add&product_id=626"},{"id":625,"sku":"13-0027","name":"Histone H3K4me2 Antibody, SNAP-Certified™ for CUT&RUN","url":"https://www.epicypher.com/products/antibodies/cutana-cut-run-antibodies/cut-run-antibodies-histone-ptms/histone-h3k4me2-antibody-snap-certified-for-cut-run","availability":"","rating":null,"brand":{"name":null},"category":["Antibodies/CUTANA™ CUT&RUN Antibodies","Antibodies/CUTANA™ CUT&RUN Antibodies/CUTANA™ CUT&RUN Antibodies to Histone PTMs","Epigenetics Kits and Reagents/CUTANA™ ChIC / CUT&RUN Assays"],"summary":"\n \n \n Type: Monoclonal\n \n \n Target Size: 15 kDa\n \n \n Format: Affi","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/625/577/Screen_Shot_2018-09-25_at_2.00.27_PM__04625.1538135192.png?c=2","alt":"Histone H3K4me2 Antibody, SNAP-Certified™ for CUT&RUN"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/625/577/Screen_Shot_2018-09-25_at_2.00.27_PM__04625.1538135192.png?c=2","alt":"Histone H3K4me2 Antibody, SNAP-Certified™ for CUT&RUN"}],"date_added":"24th Sep 2018","pre_order":false,"show_cart_action":true,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":451,"name":"Pack Size","value":"100 μg"}],"num_reviews":null,"weight":{"formatted":"0.00 LBS","value":0},"demo":false,"add_to_cart_url":"https://www.epicypher.com/cart.php?action=add&product_id=625","price":{"without_tax":{"currency":"USD","formatted":"$525.00","value":525},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=625"},{"id":628,"sku":"13-0030","name":"Histone H3K27me3 Antibody, SNAP-ChIP® Certified, CUTANA™ CUT&RUN Compatible *DISCONTINUED*","url":"https://www.epicypher.com/products/antibodies/snap-chip-certified-antibodies/histone-h3k27me3-antibody-snap-chip-certified-cutana-cut-run-compatible-discontinued","availability":"","rating":null,"brand":{"name":null},"category":[],"summary":"\n \n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/628/940/histone-h3k27me3-antibody-snap-chip-certified-cutana-cutandrun-compatible-discontinued__17318.1645734531.jpg?c=2","alt":"Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUTandRUN Compatible - DISCONTINUED"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/628/940/histone-h3k27me3-antibody-snap-chip-certified-cutana-cutandrun-compatible-discontinued__17318.1645734531.jpg?c=2","alt":"Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUTandRUN Compatible - DISCONTINUED"}],"date_added":"24th Sep 2018","pre_order":false,"show_cart_action":false,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":470,"name":"Pack Size","value":"100 μg"}],"num_reviews":null,"weight":{"formatted":"0.00 LBS","value":0},"demo":false,"add_to_wishlist_url":"/wishlist.php?action=add&product_id=628"},{"id":629,"sku":"13-0031","name":"Histone H3K36me3 Antibody, SNAP-ChIP® Certified, CUTANA™ CUT&RUN Compatible *DISCONTINUED*","url":"https://www.epicypher.com/products/nucleosomes/snap-chip-sup-sup-spike-ins/snap-chip-sup-sup-certified-antibodies/histone-h3k36me3-antibody-snap-chip-certified-cutana-cut-run-compatible-discontinued","availability":"","rating":null,"brand":{"name":null},"category":[],"summary":"\n \n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/629/581/Screen_Shot_2018-09-25_at_2.00.27_PM__10558.1538135192.png?c=2","alt":"Histone H3K36me3 Antibody, SNAP-ChIP® Certified, CUTANA™ CUT&RUN Compatible *DISCONTINUED*"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/629/581/Screen_Shot_2018-09-25_at_2.00.27_PM__10558.1538135192.png?c=2","alt":"Histone H3K36me3 Antibody, SNAP-ChIP® Certified, CUTANA™ CUT&RUN Compatible *DISCONTINUED*"}],"date_added":"24th Sep 2018","pre_order":false,"show_cart_action":false,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":446,"name":"Pack Size","value":"100 μg"}],"num_reviews":null,"weight":{"formatted":"0.00 LBS","value":0},"demo":false,"add_to_wishlist_url":"/wishlist.php?action=add&product_id=629"}],"shipping_messages":[],"rating":0,"meta_keywords":"SNAP-ChIP Barcoded Designer Nucleosomes for Chromatin IP Experiments, spike-in, chromatin IP, ChIP, chromatin immunoprecipitation, SNAP-ChIP, K-metstat, spike in, spike-in, drosophila spike-in, chip spike-in, chip spike in, chip normalization, chip-seq normalization, chip-seq spike-in, chip spike-in, chip-seq, quantitative chip spike-in, quantitative chip normalization, quantitative chip-seq normalization, quantitative chip spike-in, quantitative chip control, spike-in, drosophila spike-in, chromatin ip spike-in, chromatin ip spike in, chromatin ip normalization, chromatin ip-seq normalization, chromatin ip-seq spike-in, chromatin ip spike-in, chromatin ip-seq, quantitative chromatin ip spike-in, quantitative chromatin ip normalization, quantitative chromatin ip-seq normalization, quantitative chromatin ip spike-in, quantitative chromatin ip control, spike-in, drosophila spike-in, chromatin immunoprecipitation spike-in, chromatin immunoprecipitation spike in, chromatin immunoprecipitation normalization, chromatin immunoprecipitation-seq normalization, chromatin immunoprecipitation-seq spike-in, chromatin immunoprecipitation spike-in, chromatin immunoprecipitation-seq, quantitative chromatin immunoprecipitation spike-in, quantitative chromatin immunoprecipitation normalization, quantitative chromatin immunoprecipitation-seq normalization, quantitative chromatin immunoprecipitation spike-in, quantitative chromatin immunoprecipitation control, ChIP-seq calibration, chromatin IP calibration, ChIP calibration, chromatin immunoprecipitation calibration, ChIP-seq normalization, chromatin IP normalization, ChIP normalization, chromatin immunoprecipitation normalization, ChIP-seq spike-in, chromatin IP spike-in, ChIP spike-in, chromatin immunoprecipitation spike-in, ChIP-seq spike in, chromatin IP spike in, ChIP spike in, chromatin immunoprecipitation spike in, ChIP-seq antibody specificity, chromatin IP antibody specificity, ChIP antibody specificity, chromatin immunoprecipitation antibody specificity, ","show_quantity_input":1,"AddThisServiceButtonMeta":"","videos":{"list":[{"id":"-AlJ7Rv3beo","title_short":"An Introductio...","title_long":"An Introduction to SNAP-ChIP® Spike-in Controls","description_long":"UPDATED VIDEO: https://youtu.be/ElUS2NG5WyE.\nSNAP-ChIP (Sample Normalization and Antibody Profiling for Chromatin Immunoprecipitation) is a groundbreaking new technology that allows you to normalize your ChIP and ChIP-seq experiments as well as profile the specificity of your ChIP antibodies. Learn more about SNAP-ChIP® Spike-in Control Panels at www.epicypher.com.","description_short":"UPDATED VIDEO: https://youtu.be/ElUS2NG5WyE.\nSNAP-ChIP (Sample...","length":"03:00"}],"featured":{"id":"-AlJ7Rv3beo","title_short":"An Introductio...","title_long":"An Introduction to SNAP-ChIP® Spike-in Controls","description_long":"UPDATED VIDEO: https://youtu.be/ElUS2NG5WyE.\nSNAP-ChIP (Sample Normalization and Antibody Profiling for Chromatin Immunoprecipitation) is a groundbreaking new technology that allows you to normalize your ChIP and ChIP-seq experiments as well as profile the specificity of your ChIP antibodies. Learn more about SNAP-ChIP® Spike-in Control Panels at www.epicypher.com.","description_short":"UPDATED VIDEO: https://youtu.be/ElUS2NG5WyE.\nSNAP-ChIP (Sample...","length":"03:00"}},"title":"SNAP-ChIP® K-MetStat Panel","gift_wrapping_available":false,"min_purchase_quantity":0,"customizations":[],"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/411/282/SNAP_Chip__53961.1569012494.png?c=2","alt":"SNAP-ChIP® K-MetStat Panel"}]}
{"sku":"19-1001","url":"https://www.epicypher.com/products/nucleosomes/snap-chip-k-metstat-panel","add_this":[{"service":"facebook","annotation":""},{"service":"email","annotation":""},{"service":"print","annotation":""},{"service":"twitter","annotation":""},{"service":"linkedin","annotation":""}],"gtin":"","id":"411","bulk_discount_rates":[],"can_purchase":true,"meta_description":"SNAP-ChIP K-MetStat Panel Spike-in Normalization Control for Chromatin IP (ChIP) and ChIP-Seq","category":["Nucleosomes","Nucleosomes/SNAP-ChIP<sup>®</sup> Spike-ins"],"main_image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/411/282/SNAP_Chip__53961.1569012494.png?c=2","alt":"SNAP-ChIP® K-MetStat Panel"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=411","shipping":{"calculated":true},"num_reviews":0,"weight":"0.01 LBS","description":"<table border=\"0\" cellpadding=\"2\" cellspacing=\"2\" width=\"100%\">\n\t<tbody>\n\t\t<tr valign=\"top\">\n\t\t\t<td>\n\t\t\t<table border=\"0\" cellpadding=\"1\" cellspacing=\"1\" height=\"1490\" width=\"90%\">\n\t\t\t\t<tbody>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td valign=\"top\">\n\t\t\t\t\t\t<p></p>\n\n\t\t\t\t\t\t\n\t\t\t\t\t\t<span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><b><b>SNAP-ChIP K-MetStat Panel</b> Description:</b> A panel of distinctly modified mononucleosomes assembled from recombinant human histones expressed in <em>E. coli</em> (two each of histones H2A, H2B, H3 and H4; accession numbers: H2A-P04908; H2B-O60814; H3.1-P68431 or H3.2-Q71DI3*; H4-P62805) wrapped by 147 base pairs of barcoded Widom 601 positioning sequence DNA. The panel is comprised of a pool of 1 unmodified plus 15 histone H3 or H4 post-translational modifications (PTMs, created by a proprietary semi-synthetic method): H3K4me1, H3K4me2, H3K4me3, H3K9me1, H3K9me2, H3K9me3, H3K27me1, H3K27me2, H3K27me3, H3K36me1, H3K36me2, H3K36me3, H4K20me1, H4K20me2 and H4K20me3. Each distinctly modified nucleosome is distinguishable by a unique sequence of DNA ("barcode") at the 3' end that can be deciphered by qPCR or next-generation sequencing. Each of the 16 nucleosomes in the pool is wrapped by 2 distinct DNA species, each containing a distinct barcode ("A" and "B", see SNAP-ChIP Manual) allowing for an internal technical replicate.</span></span>\n\n\t\t\t\t\t\t<p><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><span style=\"color: orange !important;\"><b>Learn more about SNAP-ChIP Certified Antibodies <a href=\"https://www.epicypher.com/snap-chip-certified-antibodies/\">here</a></b></span></span></span></p>\n\n\t\t\t\t\t\t<p><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\">* Histone H3.2 contains a Cys to Ala mutation at position 110. </span></span><br />\n\t\t\t\t\t\t</p>\n\t\t\t\t\t\t</td>\n\t\t\t\t\t</tr>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td valign=\"top\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><b><b>SNAP-ChIP K-MetStat Panel</b> Formulation:</b> Purified recombinant mononucleosomes, containing a mixture of 16 (1 unmodified plus 15 unique) H3 and H4 PTMs in 10 mM sodium cacodylate, pH 7.5, 100 mM NaCl, 1 mM EDTA, 50% glycerol (w/v), 1x Protease Inhibitor cocktail, 100 ug/mL BSA, 10 mM β-mercaptoethanol. Molarity = 0.6 nM. MW = ~199382.1 Da (average MW of all 16 nucleosomes).</span></span><br /><br />\n\t\t\t\t\t\t</td>\n\t\t\t\t\t</tr>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td valign=\"top\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><b>SNAP-ChIP K-MetStat Panel Storage and Stability:</b> Stable for six months at -20°C from date of receipt.</span></span><br /><br />\n\t\t\t\t\t\t</td>\n\t\t\t\t\t</tr>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td valign=\"top\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><b>SNAP-ChIP K-MetStat Panel Application Notes:</b> SNAP-ChIP K-MetStats are highly purified recombinant mononucleosomes and are suitable for use as spike-in controls for ChIP reactions, for antibody specificity testing or for effector protein binding experiments. SNAP-ChIP K-MetStats from EpiCypher do not contain detectable levels of free DNA which could alter assayed activities. See manual for more information. SNAP-ChIP can easily be introduced into existing ChIP and ChIP-seq workflows for experiment normalization and monitoring of technical variability. </span></span>\n\t\t\t\t\t\t<p><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><span style=\"color: orange !important;\"><b>Request a quote <a href=\"https://www.epicypher.com/request-a-quote/\">here</a></b></span></span></span></p>\n\t\t\t\t\t\t</td>\n\t\t\t\t\t</tr>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td valign=\"top\">\n\t\t\t\t\t\t<p><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><b>Background References</b>:</span></span><br>\n\t\t\t\t\t\t <span style=\"font-family:arial,helvetica,sans-serif;\">Grzybowski A et al. 2015 Mol Cell 58: 886 - 899 <a href=\"https://www.ncbi.nlm.nih.gov/pubmed/26004229\" title=\"Calibrating ChIP-Seq with Nucleosomal Internal Standards to Measure Histone Modification Density Genome Wide\" target=\"_new\">LINK</a></span>\t\t\t\t\t\t</p>\n\t\t\t\t\t\t<p><span style=\"font-family:arial,helvetica,sans-serif;\"><span style=\"font-size:12pt;\"><b>Product References</b>:<br>\n\t\t\t\t\t\t</span></span><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\">Shah RN et al. 2018 Mol. Cell 72: 162–177 <a href=\"https://www.ncbi.nlm.nih.gov/pubmed/30244833\" title=\"Examining the Roles of H3K4 Methylation States with Systematically Characterized Antibodies\" target=\"_new\">LINK</a></span></span>\t\t\t\t\t\t<br>\n <span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\">Janssen A et al. 2018 Genes Dev. 33: 103-115 <a href=\"https://pubmed.ncbi.nlm.nih.gov/30578303/\">LINK </a><br>\n Yeganeh M et al. 2018 Nucl. Acids Res. <a href=\"https://pubmed.ncbi.nlm.nih.gov/30597109/\" title=\"Differential regulation of RNA polymerase III genes during liver regeneration\" target=\"_new\">LINK</a></span></span>\t\t\t\t\t\t<br />\n\t\t\t\t\t\t<br />\n\t\t\t\t\t </p></td>\n\t\t\t\t\t</tr>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td valign=\"top\">\n\t\t\t\t\t\t<p><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><b>SNAP-ChIP K-MetStat Panel Additional Resources and Files:</b></span></span></p>\n\n\t\t\t\t\t\t<table border=\"0\" cellpadding=\"0\" cellspacing=\"10\" height=\"447\" width=\"450\">\n\t\t\t\t\t\t\t<tbody>\n\t\t\t\t\t\t\t\t<tr>\n\t\t\t\t\t\t\t\t\t<td style=\"height:60px\" valign=\"top\" width=\"350\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Technical datasheet </strong></span></span></td>\n\t\t\t\t\t\t\t\t\t<td align=\"center\" valign=\"top\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/documents/tds/19-1001.pdf\" target=\"_new\"><img alt=\"19-1001 Datasheet\" height=\"40\" src=\"/content/documents/tds/icon.png\" width=\"30\" /></a></span></span></td>\n\t\t\t\t\t\t\t\t</tr>\n\t\t\t\t\t\t\t\t<tr>\n\t\t\t\t\t\t\t\t\t<td style=\"height:60px\" valign=\"top\" width=\"350\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>SNAP-ChIP K-MetStat Manual</strong></span></span></td>\n\t\t\t\t\t\t\t\t\t<td align=\"center\" valign=\"top\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/documents/snapchip/SNAP-ChIP_manual.pdf\" target=\"_new\"><img alt=\"SNAP-ChIP K-MetStat manual.pdf\" height=\"40\" src=\"/content/documents/tds/icon.png\" width=\"30\" /></a></span></span></td>\n\t\t\t\t\t\t\t\t</tr>\n\t\t\t\t\t\t\t\t<tr>\n\t\t\t\t\t\t\t\t\t<td style=\"height:60px\" valign=\"top\" width=\"350\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>SNAP-ChIP Brochure</strong></span></span></td>\n\t\t\t\t\t\t\t\t\t<td align=\"center\" valign=\"top\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/docs/SNAP-ChIP.pdf\" target=\"_new\"><img alt=\"SNAP-ChIP K-MetStat manual.pdf\" height=\"40\" src=\"/content/documents/tds/icon.png\" width=\"30\" /></a></span></span></td>\n\t\t\t\t\t\t\t\t</tr>\n\t\t\t\t\t\t\t\t<tr>\n\t\t\t\t\t\t\t\t\t<td style=\"height:60px\" valign=\"top\" width=\"350\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>SNAP-ChIP K-MetStat Full Panel Information: </strong>Forward/reverse barcodes, nucleosomal DNA sequences, qPCR primer sequences, TaqMan probe sequence</span></span></td>\n\t\t\t\t\t\t\t\t\t<td align=\"center\" valign=\"top\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/documents/snapchip/ SNAP-ChIP_K-MetStat_19-1001_Barcode-Sequence-Primer_Info-SHORT_ver1.1.xlsx\" target=\"_new\"><img alt=\"19-1001 Datasheet\" height=\"40\" src=\"/content/documents/excelicon.png\" width=\"30\" /></a></span></span></td>\n\t\t\t\t\t\t\t\t</tr>\n\t\t\t\t\t\t\t\t<tr>\n\t\t\t\t\t\t\t\t\t<td style=\"height:60px\" valign=\"top\" width=\"350\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>SNAP-ChIP K-MetStat qPCR Primer Sequences</strong> - Formatted for easy upload for ordering from common vendors (<em>e.g.</em> IDT)</span></span></td>\n\t\t\t\t\t\t\t\t\t<td align=\"center\" valign=\"top\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/documents/snapchip/SNAP-ChIP_K-MetStat_qPCR_Primers_BarcodeA.xlsx\" target=\"_new\"><img alt=\"19-1001 Datasheet\" height=\"40\" src=\"/content/documents/excelicon.png\" width=\"30\" /></a></span></span></td>\n\t\t\t\t\t\t\t\t</tr>\n\t\t\t\t\t\t\t\t<tr>\n\t\t\t\t\t\t\t\t\t<td style=\"height:60px\" valign=\"top\" width=\"350\"><span style=\"font-size:12pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Fasta formatted file of K-MetStat barcode Sequences</strong> - For alignment of NGS sequencing data</span></span></td>\n\t\t\t\t\t\t\t\t\t<td align=\"center\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/documents/snapchip/KmetStat_barcoded_fasta.txt.rtf\" target=\"_new\"><img alt=\"19-1001 fasta\" height=\"40\" src=\"/content/documents/excelicon.png\" width=\"30\" /></a></span></td>\n\t\t\t\t\t\t\t\t</tr>\n\t\t\t\t\t\t\t</tbody>\n\t\t\t\t\t\t</table>\n\t\t\t\t\t\t</td>\n\t\t\t\t\t</tr>\n\t\t\t\t</tbody>\n\t\t\t</table>\n\t\t\t</td>\n\t\t\t<td>\n\t\t\t<table border=\"0\" cellpadding=\"3\" cellspacing=\"3\" width=\"100%\">\n\t\t\t\t<tbody>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td align=\"center\"><span style=\"font-size:11pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/images/nucleosome/K-MetStat_Graphic.gif\" target=\"_new\"><img alt=\"K-MetStat Panel\" src=\"/content/images/nucleosome/K-MetStat_Graphic.gif\" width=\"250\" /></a></span></span></td>\n\t\t\t\t\t</tr>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td align=\"left\" valign=\"top\"><span style=\"font-size:11pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>SNAP-ChIP K-MetStat Panel:</strong> Schematic representation of nucleosomes comprising the K-MetStat Panel: H3K4me1, H3K4me2, H3K4me3, H3K9me1, H3K9me2, H3K9me3, H3K27me1, H3K27me2, H3K27me3, H3K36me1, H3K36me2, H3K36me3, H4K20me1, H4K20me2 and H4K20me3 and me0 (unmodified).<br />\n\t\t\t\t\t\t<strong>(Click image to enlarge) </strong></span></span>\n\t\t\t\t\t\t<p></p>\n\t\t\t\t\t\t</td>\n\t\t\t\t\t</tr>\n\t\t\t\t</tbody>\n\t\t\t</table>\n\n\t\t\t<table border=\"0\" cellpadding=\"3\" cellspacing=\"3\" width=\"100%\">\n\t\t\t\t<tbody>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td align=\"center\"><span style=\"font-size:11pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/images/products/nucleosomes/19-1001-DNA-gel.jpg\" target=\"_new\"><img alt=\"19-1001 DNA Gel Data\" src=\"/content/images/products/nucleosomes/19-1001-DNA-gel.jpg\" width=\"250\" /></a></span></span></td>\n\t\t\t\t\t</tr>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td align=\"left\" valign=\"top\"><span style=\"font-size:11pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>DNA Gel Data:</strong> Representative images for SNAP-ChIP K-MetStats (H3K4me0 = unmodified, H3K4me2, H3K4me3) run on a native PAGE gel and stained with ethidium bromide to visualize DNA. Lane 1: Free 147bp DNA used in nuclesome assembly (100 ng). Lane 2: Intact nucleosomes (200 ng) showing lack of free DNA. Identical experiments were performed for the entire K-MetStat Panel.<br />\n\t\t\t\t\t\t<strong>(Click image to enlarge) </strong></span></span>\n\t\t\t\t\t\t<p></p>\n\t\t\t\t\t\t</td>\n\t\t\t\t\t</tr>\n\t\t\t\t</tbody>\n\t\t\t</table>\n\n\t\t\t<table border=\"0\" cellpadding=\"3\" cellspacing=\"3\" width=\"100%\">\n\t\t\t\t<tbody>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td align=\"center\"><span style=\"font-size:11pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/images/products/nucleosomes/19-1001-protein-gel.jpg\" target=\"_new\"><img alt=\"19-1001 Protein Gel Data\" src=\"/content/images/products/nucleosomes/19-1001-protein-gel.jpg\" width=\"250\" /></a></span></span></td>\n\t\t\t\t\t</tr>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td align=\"left\" valign=\"top\"><span style=\"font-size:11pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Protein Gel Data:</strong> Representative Coomassie stained PAGE gel for SNAP-ChIP K-MetStats (2 µg each of unmodified, H3K4me1, H3K4me2, H3K4me3) to demonstrate the purity of the histones in the preparation. Sizes of molecular weight markers and positions of the core histones (H2A, H2B, H3 and H4) are indicated. Identical experiments were performed for the remainder of the K-MetStat Panel.<br />\n\t\t\t\t\t\t<strong>(Click image to enlarge) </strong></span></span>\n\t\t\t\t\t\t<p></p>\n\t\t\t\t\t\t</td>\n\t\t\t\t\t</tr>\n\t\t\t\t</tbody>\n\t\t\t</table>\n\n\t\t\t<table border=\"0\" cellpadding=\"3\" cellspacing=\"3\" width=\"100%\">\n\t\t\t\t<tbody>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td align=\"center\"><span style=\"font-size:11pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"/content/images/products/nucleosomes/19-1001-ChIP-data.jpg\" target=\"_new\"><img alt=\"19-1001 ChIP Data\" src=\"/content/images/products/nucleosomes/19-1001-ChIP-data.jpg\" width=\"250\" /></a></span></span></td>\n\t\t\t\t\t</tr>\n\t\t\t\t\t<tr>\n\t\t\t\t\t\t<td align=\"left\" valign=\"top\">\n\t\t\t\t\t\t<p><span style=\"font-size:11pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>ChIP Data:</strong> Representative images for SNAP-ChIP K-MetStats (unmodified, H3K4me1, H3K4me2, H3K4me3) assayed in a chromatin immunoprecipitation (ChIP) experiment using commercially available ChIP grade antibodies (3 µg, n = 3). Quantitative Real-Time PCR (qPCR) for the DNA barcodes corresponding to unmodified (H3K4me0), H3K4me1, H3K4me2, and H3K4me3 nucleosomes show recovery of the barcodes corresponding to the expected antibody target. Identical experiments were performed for the remainder of the K-MetStat Panel (H3K9me1, H3K9me2, H3K9me3, H3K27me1, H3K27me2, H3K27me3, H3K36me1, H3K36me2, H3K36me3, H4K20me1, H4K20me2 and H4K20me3).</span></span></p>\n\n\t\t\t\t\t\t<p><span style=\"font-size:11pt;\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>N.B.</strong>-SNAP-ChIP is extremely useful for determining antibody cross-reactivity with off-target modifications. The cross-reactivity observed above (K4me1 & K4me2) is not indicative of a failure in the SNAP-ChIP procedure, but in the antibody to specifically recognize its target.<br />\n\t\t\t\t\t\t<strong>(Click image to enlarge) </strong></span></span></p>\n\n\t\t\t\t\t\t<p></p>\n\t\t\t\t\t\t</td>\n\t\t\t\t\t</tr>\n\t\t\t\t</tbody>\n\t\t\t</table>\n\n\t\t\t<table border=\"0\" cellpadding=\"3\" cellspacing=\"3\" width=\"100%\">\n\t\t\t\t<tbody>\n\t\t\t\t</tbody>\n\t\t\t</table>\n\t\t\t</td>\n\t\t</tr>\n\t</tbody>\n</table>\n\n","tags":[],"warranty":"","price":{"without_tax":{"formatted":"$365.00","value":365,"currency":"USD"},"tax_label":"Sales Tax"},"detail_messages":"","availability":"","page_title":"SNAP-ChIP K-MetStat Panel Spike-in Normalization Control for Chromatin IP (ChIP) and ChIP-Seq","cart_url":"https://www.epicypher.com/cart.php","max_purchase_quantity":0,"mpn":"","upc":null,"options":[{"id":682,"type":"Configurable_PickList_Set","display_name":"Select Pack Size:","required":true,"condition":true,"state":"variant_option","values":[{"label":"20 µl","id":111,"data":"20 µl","selected":true},{"label":"200 µl","id":106,"data":"200 µl","selected":false}],"partial":"set-radio"}],"related_products":[{"id":555,"sku":null,"name":"SNAP-ChIP® OncoStat Panel","url":"https://www.epicypher.com/products/nucleosomes/snap-chip-oncostat-panel","availability":"","rating":null,"brand":{"name":null},"category":["Nucleosomes","Nucleosomes/SNAP-ChIP<sup>®</sup> Spike-ins"],"summary":" \n\n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/555/399/OncoStat__15594.1569012525.png?c=2","alt":"SNAP-ChIP® OncoStat Panel"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/555/399/OncoStat__15594.1569012525.png?c=2","alt":"SNAP-ChIP® OncoStat Panel"}],"date_added":"31st Jul 2018","pre_order":false,"show_cart_action":true,"has_options":true,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":null,"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"price":{"without_tax":{"currency":"USD","formatted":"$365.00","value":365},"price_range":{"min":{"without_tax":{"currency":"USD","formatted":"$365.00","value":365},"tax_label":"Sales Tax"},"max":{"without_tax":{"currency":"USD","formatted":"$2,795.00","value":2795},"tax_label":"Sales Tax"}},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=555"},{"id":626,"sku":"13-0028","name":"Histone H3K4me3 Antibody, SNAP-ChIP Certified *DISCONTINUED*","url":"https://www.epicypher.com/products/antibodies/snap-chip-certified-antibodies/histone-h3k4me3-antibody-snap-chip-certified","availability":"","rating":null,"brand":{"name":null},"category":[],"summary":"\n \n \n \n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/626/578/Screen_Shot_2018-09-25_at_2.00.27_PM__52796.1538135192.png?c=2","alt":"Histone H3K4me3 Antibody, SNAP-ChIP Certified *DISCONTINUED*"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/626/578/Screen_Shot_2018-09-25_at_2.00.27_PM__52796.1538135192.png?c=2","alt":"Histone H3K4me3 Antibody, SNAP-ChIP Certified *DISCONTINUED*"}],"date_added":"24th Sep 2018","pre_order":false,"show_cart_action":false,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":452,"name":"Pack Size","value":"100 μg"}],"num_reviews":null,"weight":{"formatted":"0.00 LBS","value":0},"demo":false,"add_to_wishlist_url":"/wishlist.php?action=add&product_id=626"},{"id":625,"sku":"13-0027","name":"Histone H3K4me2 Antibody, SNAP-Certified™ for CUT&RUN","url":"https://www.epicypher.com/products/antibodies/cutana-cut-run-antibodies/cut-run-antibodies-histone-ptms/histone-h3k4me2-antibody-snap-certified-for-cut-run","availability":"","rating":null,"brand":{"name":null},"category":["Antibodies/CUTANA™ CUT&RUN Antibodies","Antibodies/CUTANA™ CUT&RUN Antibodies/CUTANA™ CUT&RUN Antibodies to Histone PTMs","Epigenetics Kits and Reagents/CUTANA™ ChIC / CUT&RUN Assays"],"summary":"\n \n \n Type: Monoclonal\n \n \n Target Size: 15 kDa\n \n \n Format: Affi","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/625/577/Screen_Shot_2018-09-25_at_2.00.27_PM__04625.1538135192.png?c=2","alt":"Histone H3K4me2 Antibody, SNAP-Certified™ for CUT&RUN"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/625/577/Screen_Shot_2018-09-25_at_2.00.27_PM__04625.1538135192.png?c=2","alt":"Histone H3K4me2 Antibody, SNAP-Certified™ for CUT&RUN"}],"date_added":"24th Sep 2018","pre_order":false,"show_cart_action":true,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":451,"name":"Pack Size","value":"100 μg"}],"num_reviews":null,"weight":{"formatted":"0.00 LBS","value":0},"demo":false,"add_to_cart_url":"https://www.epicypher.com/cart.php?action=add&product_id=625","price":{"without_tax":{"currency":"USD","formatted":"$525.00","value":525},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=625"},{"id":628,"sku":"13-0030","name":"Histone H3K27me3 Antibody, SNAP-ChIP® Certified, CUTANA™ CUT&RUN Compatible *DISCONTINUED*","url":"https://www.epicypher.com/products/antibodies/snap-chip-certified-antibodies/histone-h3k27me3-antibody-snap-chip-certified-cutana-cut-run-compatible-discontinued","availability":"","rating":null,"brand":{"name":null},"category":[],"summary":"\n \n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/628/940/histone-h3k27me3-antibody-snap-chip-certified-cutana-cutandrun-compatible-discontinued__17318.1645734531.jpg?c=2","alt":"Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUTandRUN Compatible - DISCONTINUED"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/628/940/histone-h3k27me3-antibody-snap-chip-certified-cutana-cutandrun-compatible-discontinued__17318.1645734531.jpg?c=2","alt":"Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUTandRUN Compatible - DISCONTINUED"}],"date_added":"24th Sep 2018","pre_order":false,"show_cart_action":false,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":470,"name":"Pack Size","value":"100 μg"}],"num_reviews":null,"weight":{"formatted":"0.00 LBS","value":0},"demo":false,"add_to_wishlist_url":"/wishlist.php?action=add&product_id=628"},{"id":629,"sku":"13-0031","name":"Histone H3K36me3 Antibody, SNAP-ChIP® Certified, CUTANA™ CUT&RUN Compatible *DISCONTINUED*","url":"https://www.epicypher.com/products/nucleosomes/snap-chip-sup-sup-spike-ins/snap-chip-sup-sup-certified-antibodies/histone-h3k36me3-antibody-snap-chip-certified-cutana-cut-run-compatible-discontinued","availability":"","rating":null,"brand":{"name":null},"category":[],"summary":"\n \n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/629/581/Screen_Shot_2018-09-25_at_2.00.27_PM__10558.1538135192.png?c=2","alt":"Histone H3K36me3 Antibody, SNAP-ChIP® Certified, CUTANA™ CUT&RUN Compatible *DISCONTINUED*"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/629/581/Screen_Shot_2018-09-25_at_2.00.27_PM__10558.1538135192.png?c=2","alt":"Histone H3K36me3 Antibody, SNAP-ChIP® Certified, CUTANA™ CUT&RUN Compatible *DISCONTINUED*"}],"date_added":"24th Sep 2018","pre_order":false,"show_cart_action":false,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":446,"name":"Pack Size","value":"100 μg"}],"num_reviews":null,"weight":{"formatted":"0.00 LBS","value":0},"demo":false,"add_to_wishlist_url":"/wishlist.php?action=add&product_id=629"}],"shipping_messages":[],"rating":0,"meta_keywords":"SNAP-ChIP Barcoded Designer Nucleosomes for Chromatin IP Experiments, spike-in, chromatin IP, ChIP, chromatin immunoprecipitation, SNAP-ChIP, K-metstat, spike in, spike-in, drosophila spike-in, chip spike-in, chip spike in, chip normalization, chip-seq normalization, chip-seq spike-in, chip spike-in, chip-seq, quantitative chip spike-in, quantitative chip normalization, quantitative chip-seq normalization, quantitative chip spike-in, quantitative chip control, spike-in, drosophila spike-in, chromatin ip spike-in, chromatin ip spike in, chromatin ip normalization, chromatin ip-seq normalization, chromatin ip-seq spike-in, chromatin ip spike-in, chromatin ip-seq, quantitative chromatin ip spike-in, quantitative chromatin ip normalization, quantitative chromatin ip-seq normalization, quantitative chromatin ip spike-in, quantitative chromatin ip control, spike-in, drosophila spike-in, chromatin immunoprecipitation spike-in, chromatin immunoprecipitation spike in, chromatin immunoprecipitation normalization, chromatin immunoprecipitation-seq normalization, chromatin immunoprecipitation-seq spike-in, chromatin immunoprecipitation spike-in, chromatin immunoprecipitation-seq, quantitative chromatin immunoprecipitation spike-in, quantitative chromatin immunoprecipitation normalization, quantitative chromatin immunoprecipitation-seq normalization, quantitative chromatin immunoprecipitation spike-in, quantitative chromatin immunoprecipitation control, ChIP-seq calibration, chromatin IP calibration, ChIP calibration, chromatin immunoprecipitation calibration, ChIP-seq normalization, chromatin IP normalization, ChIP normalization, chromatin immunoprecipitation normalization, ChIP-seq spike-in, chromatin IP spike-in, ChIP spike-in, chromatin immunoprecipitation spike-in, ChIP-seq spike in, chromatin IP spike in, ChIP spike in, chromatin immunoprecipitation spike in, ChIP-seq antibody specificity, chromatin IP antibody specificity, ChIP antibody specificity, chromatin immunoprecipitation antibody specificity, ","show_quantity_input":1,"AddThisServiceButtonMeta":"","videos":{"list":[{"id":"-AlJ7Rv3beo","title_short":"An Introductio...","title_long":"An Introduction to SNAP-ChIP® Spike-in Controls","description_long":"UPDATED VIDEO: https://youtu.be/ElUS2NG5WyE.\nSNAP-ChIP (Sample Normalization and Antibody Profiling for Chromatin Immunoprecipitation) is a groundbreaking new technology that allows you to normalize your ChIP and ChIP-seq experiments as well as profile the specificity of your ChIP antibodies. Learn more about SNAP-ChIP® Spike-in Control Panels at www.epicypher.com.","description_short":"UPDATED VIDEO: https://youtu.be/ElUS2NG5WyE.\nSNAP-ChIP (Sample...","length":"03:00"}],"featured":{"id":"-AlJ7Rv3beo","title_short":"An Introductio...","title_long":"An Introduction to SNAP-ChIP® Spike-in Controls","description_long":"UPDATED VIDEO: https://youtu.be/ElUS2NG5WyE.\nSNAP-ChIP (Sample Normalization and Antibody Profiling for Chromatin Immunoprecipitation) is a groundbreaking new technology that allows you to normalize your ChIP and ChIP-seq experiments as well as profile the specificity of your ChIP antibodies. Learn more about SNAP-ChIP® Spike-in Control Panels at www.epicypher.com.","description_short":"UPDATED VIDEO: https://youtu.be/ElUS2NG5WyE.\nSNAP-ChIP (Sample...","length":"03:00"}},"title":"SNAP-ChIP® K-MetStat Panel","gift_wrapping_available":false,"min_purchase_quantity":0,"customizations":[],"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/411/282/SNAP_Chip__53961.1569012494.png?c=2","alt":"SNAP-ChIP® K-MetStat Panel"}]}
|
|
Current stock:
0