CUTANA™ ChIC/CUT&RUN Kit
{"url":"https://www.epicypher.com/products/epigenetics-reagents-and-assays/cutana-chic-cut-and-run-kit","add_this":[{"service":"facebook","annotation":""},{"service":"email","annotation":""},{"service":"print","annotation":""},{"service":"twitter","annotation":""},{"service":"linkedin","annotation":""}],"gtin":"","id":"772","bulk_discount_rates":[],"can_purchase":true,"meta_description":"CUTANA™ ChIC/CUT&RUN Kit includes all needed reagents to go from cells to purified CUT&RUN DNA. ","category":["Epigenetics Kits and Reagents","Epigenetics Kits and Reagents/CUTANA™ ChIC / CUT&RUN Assays"],"AddThisServiceButtonMeta":"","main_image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/772/998/Epicypher_CUTANA_CUTRUN_Box__31908.1646406725.png?c=2","alt":"CUTANA™ ChIC/CUT&RUN Kit"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=772","shipping":{"calculated":true},"num_reviews":0,"weight":"0.01 LBS","custom_fields":[{"id":"588","name":"Pack size","value":"48 Reactions"}],"sku":"14-1048","description":"<div class=\"product-general-info\">\n <ul style=\"display: none\" class=\"product-general-info__list-left\">\n <li class=\"product-general-info__list-item\"></li>\n <li class=\"product-general-info__list-item\"></li>\n </ul>\n <ul style=\"display: none\" class=\"product-general-info__list-right\">\n <li class=\"product-general-info__list-item\"></li>\n <li class=\"product-general-info__list-item\"></li>\n </ul>\n <ul style=\"padding: 0\" class=\"product-general-info__list-right\">\n <li class=\"product-general-info__list-item\">\n <a href=\"#bioz\">\n <div\n id=\"w-s-3835-14-1048\"\n style=\"\n width: 240px;\n height: 58px;\n position: relative;\n overflow-y: hidden;\n \"\n ></div>\n <div id=\"bioz-w-pb-14-1048-div\">\n <a\n id=\"bioz-w-pb-14-1048\"\n style=\"font-size: 12px; color: transparent\"\n href=\"https://www.bioz.com/\"\n target=\"_blank\"\n >\n <img\n src=\"https://cdn.bioz.com/assets/favicon.png\"\n style=\"\n width: 11px;\n height: 11px;\n vertical-align: baseline;\n padding-bottom: 0px;\n margin-left: 0px;\n margin-bottom: 0px;\n float: none;\n display: none;\n \"\n />\n </a></div\n ></a>\n </li>\n </ul>\n</div>\n<div class=\"service_accordion product-droppdown\">\n <div class=\"container\">\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel current\">\n <h3 class=\"sub-title1\">Description</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description-specific\"\n >\n <p>\n The CUTANA™ ChIC/CUT&RUN Kit enables streamlined chromatin\n profiling of histone post-translational modifications (PTMs)\n and chromatin associated proteins.\n </p>\n <p\n style=\"\n background-color: #4698cb;\n color: #fff;\n padding: 1.3rem;\n text-align: center;\n border-radius: 12px;\n \"\n >\n <a\n target=\"_blank\"\n style=\"color: #fff\"\n href=\"/products/epigenetics-reagents-and-assays/cutana-cut-run-library-prep-kit\"\n >Save 10% when you buy with our Library Prep Kit for a\n streamlined CUT&RUN workflow! Learn more >>\n </a>\n </p>\n <p>\n The CUT&RUN Kit Version 4 (v4) now uses SPRI magnetic beads\n for DNA purification instead of DNA spin columns, enabling\n multi-channel pipetting throughout for increased throughput\n and reproducibility. Positive (H3K4me3) and negative (IgG)\n control antibodies are included to pair with SNAP-CUTANA™\n spike-in controls for assay optimization and continuous assay\n monitoring (<strong>Figure 2</strong>). <i>E. coli</i> DNA is\n included for data normalization. The kit is compatible with a\n variety of inputs including cells or nuclei derived from\n native, cryopreserved, or cross-linked samples. While it is\n recommended to start with 500,000 cells, comparable data can\n be generated using as few as 5,000 cells. The inclusion of\n controls, as well as compatibility with diverse target types,\n sample inputs, and low cell numbers, make this kit the go-to\n solution for chromatin mapping experiments. To place bulk\n orders, contact\n <a href=\"mailto:info@epicypher.com\">info@epicypher.com</a>.\n </p>\n <p>\n Keep learning by checking out our\n <a href=\"https://support.epicypher.com/\" target=\"_blank\"\n >Tech Support Center</a\n >.\n </p>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel current\">\n <h3 class=\"sub-title1\">Validation Data</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description-specific\"\n >\n <section class=\"image-picker\">\n <div class=\"image-picker__left\">\n <div\n class=\"image-picker__main-content_active image-picker__main-content\"\n >\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg\n class=\"image-picker__svg-left\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\"\n >\n <path\n d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\"\n />\n </svg>\n </button>\n <a\n href=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/figure1.jpg?t=1699317893\"\n target=\"_new\"\n class=\"image-picker__main-image-link\"\n ><img\n loading=\"lazy\"\n alt=\"14-1048-DFSDA\"\n src=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/figure1.jpg?t=1699317893\"\n loading=\"lazy\"\n class=\"image-picker__main-image\"\n />\n <span class=\"image-picker__main-image-caption\"\n >(Click to enlarge)</span\n ></a\n >\n <button class=\"image-picker__right-arrow\">\n <svg\n class=\"image-picker__svg-right\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\"\n >\n <path\n d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\"\n />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\"\n ><strong\n >Figure 1: CUT&RUN DNA fragment size distribution\n analysis</strong\n ><br />\n CUT&RUN was performed as described in\n <strong>Figure 5</strong>. Library DNA was analyzed by\n Agilent Tapestation<sup>®</sup>. This analysis\n confirmed that mononucleosomes were predominantly\n enriched in CUT&RUN (~300 bp peaks represent 150 bp\n nucleosomes + sequencing adapters).\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg\n class=\"image-picker__svg-left\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\"\n >\n <path\n d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\"\n />\n </svg>\n </button>\n <a\n href=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/14-1048-23298006-81-figure-2.jpg?t=1698650317\"\n target=\"_new\"\n class=\"image-picker__main-image-link\"\n ><img\n loading=\"lazy\"\n alt=\"14-1048-k-metstat-spike-in-controls\"\n src=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/14-1048-23298006-81-figure-2.jpg?t=1698650317\"\n class=\"image-picker__main-image\"\n loading=\"lazy\"\n />\n <span class=\"image-picker__main-image-caption\"\n >(Click to enlarge)</span\n ></a\n >\n <button class=\"image-picker__right-arrow\">\n <svg\n class=\"image-picker__svg-right\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\"\n >\n <path\n d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\"\n />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\"\n ><strong\n >Figure 2: SNAP-CUTANA™ K-MetStat Spike-in\n controls</strong\n ><br />\n DNA-barcoded designer nucleosomes (dNucs) representing\n 16 K-methyl PTMs: mono-, di-, and tri-methylation at\n H3K4, H3K9, H3K27, H3K36, and H4K20, as well as\n unmodified control, were spiked into CUT&RUN reactions\n prior to the addition of antibodies (IgG, H3K4me3).\n Instances of each spike-in barcode were counted and\n normalized from raw fastq files using the shell script\n and analysis excel sheet available on the spike-in\n product page (EpiCypher\n <a\n href=\"https://www.epicypher.com/products/nucleosomes/snap-cutana-k-metstat-panel\"\n target=\"_blank\"\n >19-1002</a\n >). Barcodes for IgG (top; normalized to total reads)\n and H3K4me3 (bottom; normalized to on-target)\n antibodies are shown. The spike-ins confirmed optimal\n experimental conditions (H3K4me3 antibody specifically\n recovered the target dNuc, while IgG showed no\n preferential enrichment).\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg\n class=\"image-picker__svg-left\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\"\n >\n <path\n d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\"\n />\n </svg>\n </button>\n <a\n href=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/14-1048-23298006-81-figure-3.jpg?t=1698693349\"\n target=\"_new\"\n class=\"image-picker__main-image-link\"\n >\n <img\n loading=\"lazy\"\n alt=\"14-1048-CRGWH\"\n src=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/14-1048-23298006-81-figure-3.jpg?t=1698693349\"\n loading=\"lazy\"\n class=\"image-picker__main-image\"\n />\n <span class=\"image-picker__main-image-caption\"\n >(Click to enlarge)</span\n >\n </a>\n <button class=\"image-picker__right-arrow\">\n <svg\n class=\"image-picker__svg-right\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\"\n >\n <path\n d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\"\n />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\">\n <strong>Figure 3: CUT&RUN genome-wide heatmaps</strong\n ><br />\n CUT&RUN was performed as described in\n <strong>Figure 5</strong>. Heatmaps show two\n replicates (“Rep”) of IgG (left) and H3K4me3 (right)\n kit control antibodies in aligned rows ranked by\n intensity (top to bottom) and colored such that red\n indicates high localized enrichment and blue denotes\n background signal.\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg\n class=\"image-picker__svg-left\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\"\n >\n <path\n d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\"\n />\n </svg>\n </button>\n <a\n href=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/14-1048-23298006-81-figure-4.jpg?t=1698693353\"\n target=\"_new\"\n class=\"image-picker__main-image-link\"\n >\n <img\n loading=\"lazy\"\n alt=\"14-1048-RGBT\"\n src=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/14-1048-23298006-81-figure-4.jpg?t=1698693353\"\n loading=\"lazy\"\n class=\"image-picker__main-image\"\n />\n <span class=\"image-picker__main-image-caption\"\n >(Click to enlarge)</span\n >\n </a>\n <button class=\"image-picker__right-arrow\">\n <svg\n class=\"image-picker__svg-right\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\"\n >\n <path\n d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\"\n />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\">\n <strong\n >Figure 4: Representative gene browser\n tracks</strong\n ><br />\n CUT&RUN was performed as described in\n <strong>Figure 5</strong>. A representative 174 kb\n window at the TRMT2A gene is shown for two replicates\n (“Rep”) of IgG and H3K4me3 kit control antibodies.\n Representative tracks are also shown for antibodies to\n H3K27me3 and the transcription factor CTCF. The\n CUT&RUN kit produced the expected genomic distribution\n for each target. Images were generated using the\n Integrative Genomics Viewer (IGV, Broad Institute).\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg\n class=\"image-picker__svg-left\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\"\n >\n <path\n d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\"\n />\n </svg>\n </button>\n <a\n href=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/14-1048-23298006-81-figure-5.png?t=1698693354\"\n target=\"_new\"\n class=\"image-picker__main-image-link\"\n >\n <img\n loading=\"lazy\"\n alt=\"14-1048-RGBT\"\n src=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/14-1048-23298006-81-figure-5.png?t=1698693354\"\n loading=\"lazy\"\n class=\"image-picker__main-image\"\n />\n <span class=\"image-picker__main-image-caption\"\n >(Click to enlarge)</span\n >\n </a>\n <button class=\"image-picker__right-arrow\">\n <svg\n class=\"image-picker__svg-right\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\"\n >\n <path\n d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\"\n />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\">\n <strong>Figure 5: CUT&RUN methods</strong><br />\n CUT&RUN was performed using the CUTANA™ ChIC/CUT&RUN\n Kit starting with 500k K562 cells with 0.5 µg of\n either IgG (EpiCypher\n <a\n href=\"https://www.epicypher.com/products/nucleosomes/snap-cutana-spike-in-controls/cutana-rabbit-igg-cut-run-negative-control-antibody\"\n target=\"_blank\"\n >13-0042</a\n >), H3K4me3 (EpiCypher\n <a\n href=\"https://www.epicypher.com/products/antibodies/snap-chip-certified-antibodies/histone-h3k4me3-antibody-snap-chip-certified-cutana-cut-run-compatible\"\n target=\"_blank\"\n >13-0041</a\n >), H3K27me3 (EpiCypher\n <a\n href=\"https://www.epicypher.com/products/antibodies/h3k27me3-antibody-snap-certified-for-cut-run-and-cut-and-tag\"\n target=\"_blank\"\n >13-0055</a\n >), or 0.125 µg of CTCF (EpiCypher\n <a\n href=\"https://www.epicypher.com/products/antibodies/cutana-cut-run-compatible-antibodies/ctcf-cutana-cut-and-run-antibody\"\n target=\"_blank\"\n >13-2014</a\n >) antibodies in duplicate. Library preparation was\n performed with 5 ng of DNA (or the total amount\n recovered if less than 5 ng) using the CUTANA™ Library\n Prep Kit (EpiCypher\n <a\n href=\"https://www.epicypher.com/products/epigenetics-reagents-and-assays/cutana-cut-and-run-library-prep-kit\"\n target=\"_blank\"\n >14-1001/14-1002</a\n >). Libraries were run on an Illumina NextSeq2000 with\n paired-end sequencing (2x50 bp). Sample sequencing\n depth was 3.5 million reads (IgG Rep 1), 3.8 million\n reads (IgG Rep 2), 4.7 million reads (H3K4me3 Rep 1),\n 6.9 million reads (H3K4me3 Rep 2), 6.6 million reads\n (H3K27me3 Rep 1), 4.7 million reads (H3K27me3 Rep 2),\n 3.9 million reads (CTCF Rep 1) and 4.6 million reads\n (CTCF Rep 2). Data were aligned to the hg19 genome\n using Bowtie2. Data were filtered to remove\n duplicates, multi-aligned reads, and ENCODE DAC Exclusion List\n regions.\n </span>\n </p>\n </div>\n </div>\n <aside class=\"image-picker__right\">\n <div class=\"image-picker__gallery\">\n <img\n loading=\"lazy\"\n alt=\"14-1048-DFSDA\"\n src=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/figure1.jpg?t=1699317893\"\n width=\"200\"\n loading=\"lazy\"\n class=\"image-picker__side-image image-picker__side-image_active\"\n role=\"button\"\n />\n <img\n loading=\"lazy\"\n alt=\"14-1048-k-metstat-spike-in-controls\"\n src=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/14-1048-23298006-81-figure-2.jpg?t=1698650317\"\n class=\"image-picker__side-image\"\n loading=\"lazy\"\n role=\"button\"\n />\n <img\n loading=\"lazy\"\n alt=\"14-1048-CRGWH\"\n src=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/14-1048-23298006-81-figure-3.jpg?t=1698693349\"\n loading=\"lazy\"\n class=\"image-picker__side-image\"\n role=\"button\"\n />\n <img\n loading=\"lazy\"\n alt=\"14-1048-RGBT\"\n src=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/14-1048-23298006-81-figure-4.jpg?t=1698693353\"\n loading=\"lazy\"\n class=\"image-picker__side-image\"\n role=\"button\"\n />\n <img\n loading=\"lazy\"\n alt=\"14-1048-RGBT\"\n src=\"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/original/image-manager/14-1048-23298006-81-figure-5.png?t=1698693354\"\n loading=\"lazy\"\n class=\"image-picker__side-image\"\n role=\"button\"\n />\n </div>\n </aside>\n </section>\n </div>\n </div>\n </div>\n\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Kit Contents</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\"\n >\n <table class=\"epicypher-table\">\n <tr>\n <th>Item</th>\n <th>Cat. No.</th>\n </tr>\n\n <tr>\n <td>CUTANA™ pAG-MNase for ChIC/CUT&RUN Workflows</td>\n <td>\n <a\n href=\"https://www.epicypher.com/products/epigenetics-reagents-and-assays/cutana-pag-mnase-for-chic-cut-and-run-workflows\"\n >15-1016</a\n >\n </td>\n </tr>\n\n <tr>\n <td>H3K4me3 Antibody, SNAP-Certified™ for CUT&RUN</td>\n <td>\n <a\n href=\"https://www.epicypher.com/products/antibodies/snap-chip-certified-antibodies/histone-h3k4me3-antibody-snap-chip-certified-cutana-cut-run-compatible\"\n >13-0041</a\n >k\n </td>\n </tr>\n\n <tr>\n <td>\n CUTANA™ Rabbit IgG CUT&RUN Negative Control Antibody\n </td>\n <td>\n <a\n href=\"https://www.epicypher.com/products/nucleosomes/snap-cutana-spike-in-controls/cutana-rabbit-igg-cut-run-negative-control-antibody\"\n >13-0042</a\n >k\n </td>\n </tr>\n\n <tr>\n <td>SNAP-CUTANA™ K-MetStat Panel</td>\n <td>\n <a\n href=\"https://www.epicypher.com/products/nucleosomes/snap-cutana-k-metstat-panel\"\n >19-1002</a\n >k\n </td>\n </tr>\n\n <tr>\n <td>\n CUTANA™ Concanavalin A Conjugated Paramagnetic Beads\n </td>\n <td>\n <a\n href=\"https://www.epicypher.com/products/epigenetics-kits-and-reagents/cutana-chic-cut-run-assays/cutana-concanavalin-a-conjugated-paramagnetic-beads\"\n >21-1401</a\n >\n </td>\n </tr>\n\n <tr>\n <td>CUTANA™ <i>E. coli</i> Spike-in DNA</td>\n <td>\n <a\n href=\"https://www.epicypher.com/products/nucleosomes/snap-cutana-spike-in-controls/cutana-e-coli-spike-in-dna\"\n >18-1401</a\n >\n </td>\n </tr>\n\n <tr>\n <td>CUTANA™ CUT&RUN 8-Strip 0.2 mL Tubes</td>\n <td>\n <a\n href=\"https://www.epicypher.com/products/epigenetics-kits-and-reagents/cutana-chic-cut-run-assays/cutana-cut-run-8-strip-0-2-ml-tubes\"\n >10-0009</a\n >a\n </td>\n </tr>\n\n <tr>\n <td>Bead Activation Buffer</td>\n <td>21-1001</td>\n </tr>\n\n <tr>\n <td>Pre-Wash Buffer</td>\n <td>21-1002</td>\n </tr>\n\n <tr>\n <td>Stop Buffer</td>\n <td>21-1003</td>\n </tr>\n\n <tr>\n <td>5% Digitonin</td>\n <td>21-1004k</td>\n </tr>\n <tr>\n <td>1 M Spermidine</td>\n <td>21-1005</td>\n </tr>\n <tr>\n <td>0.5 M EDTA</td>\n <td>21-1006</td>\n </tr>\n <tr>\n <td>100 mM Calcium Chloride</td>\n <td>21-1007</td>\n </tr>\n <tr>\n <td>0.1X TE Buffer</td>\n <td>21-1025</td>\n </tr>\n <tr>\n <td>\n SPRIselect reagent manufactured by Beckman Coulter, Inc.\n </td>\n <td>21-1405</td>\n </tr>\n </table>\n </div>\n </div>\n </div>\n\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Recommended Accessory Products</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\"\n >\n <table class=\"epicypher-table\">\n <tr>\n <th>Item</th>\n <th>Cat. No.</th>\n </tr>\n\n <tr>\n <td>CUTANA™ CUT&RUN Library Prep Kit</td>\n <td>\n <a\n href=\"/products/epigenetics-reagents-and-assays/cutana-cut-and-run-library-prep-kit\"\n >14-1001/14-1002</a\n >\n </td>\n </tr>\n\n <tr>\n <td>SNAP-CUTANA™ K-MetStat Panel</td>\n <td>\n <a\n href=\"/products/nucleosomes/snap-cutana-k-metstat-panel\"\n >19-1002</a\n >\n </td>\n </tr>\n\n <tr>\n <td id=\"specific-row\">CUT&RUN Antibodies</td>\n <td>\n <a\n href=\"/products/antibodies/cutana-cut-and-run-antibodies\"\n >See the list</a\n >\n </td>\n </tr>\n\n <tr>\n <td id=\"specific-row\">\n Magnetic Separation Rack, 0.2 mL Tubes\n </td>\n <td>\n <a\n href=\"/products/epigenetics-reagents-and-assays/cutana-chic-cut-run-assays/magnetic-separation-rack-0-2-ml-tubes\"\n >10-0008</a\n >\n </td>\n </tr>\n\n <tr>\n <td id=\"specific-row\">\n Magnetic Separation Rack, 1.5 mL Tubes\n </td>\n <td>\n <a\n href=\"/products/epigenetics-reagents-and-assays/magnetic-separation-rack-1-5-ml-tubes\"\n >10-0012</a\n >\n </td>\n </tr>\n <tr>\n <td id=\"specific-row\">CUTANA™ Nuclei Extraction Buffer</td>\n <td>\n <a\n href=\"https://www.epicypher.com/products.php?product=CUTANA%E2%84%A2-Nuclei-Extraction-Buffer&showHidden=true\"\n >21-1026</a\n >\n </td>\n </tr>\n </table>\n </div>\n </div>\n </div>\n\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Technical Information</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\"\n >\n <div class=\"product-tech-info\">\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Storage</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n OPEN KIT IMMEDIATELY and store components at room\n temperature, 4°C, and -20°C as indicated (see\n <strong>User Manual corresponding to Kit Version 4</strong\n >). Stable for 6 months upon date of receipt.\n </div>\n </div>\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Instructions for Use</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n See User Manual corresponding to Kit Version 4. This kit\n is not compatible with previous user manuals.\n <strong\n >Version 4 contains SPRI magnetic beads for DNA clean\n up, while previous versions used DNA spin\n columns.</strong\n >.\n </div>\n </div>\n </div>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Additional Information</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\"\n >\n <p>\n Beckman Coulter, the stylized logo, and SPRIselect are\n trademarks or registered trademarks of Beckman Coulter, Inc.\n in the United States and other countries.\n </p>\n </div>\n </div>\n </div>\n\n <h3\n style=\"color: #4698cb; font-size: 24px; border-bottom: none\"\n class=\"custom-title-description\"\n >\n Documents & Resources\n </h3>\n\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <!-- <h3 style=\"color: #4698cb; margin-bottom: 0\">Current Lot</h3> -->\n <h3 style=\"margin-top: 1rem; padding-top: 0\" class=\"sub-title1\">\n Kit Version 4\n </h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\"\n >\n <div class=\"product-documents\">\n <a\n href=\"/content/documents/tds/14-1048.pdf\"\n target=\"_new\"\n class=\"product-documents__link\"\n >\n <svg\n version=\"1.1\"\n id=\"Layer_1\"\n xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\"\n x=\"0px\"\n y=\"0px\"\n viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\"\n xml:space=\"preserve\"\n class=\"product-documents__icon\"\n alt=\"16-0002 Datasheet\"\n >\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M191.92,68.77l-47.69-47.69c-1.33-1.33-3.12-2.08-5.01-2.08H45.09C41.17,19,38,22.17,38,26.09v184.36\n c0,3.92,3.17,7.09,7.09,7.09h141.82c3.92,0,7.09-3.17,7.09-7.09V73.8C194,71.92,193.25,70.1,191.92,68.77z M177.65,77.06h-41.7\n v-41.7L177.65,77.06z M178.05,201.59H53.95V34.95h66.92v47.86c0,5.14,4.17,9.31,9.31,9.31h47.86V201.59z\"\n />\n </g>\n <rect\n x=\"20\"\n y=\"112\"\n class=\"product-documents__svg-background\"\n width=\"146\"\n height=\"76\"\n />\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M23.83,125.68h22.36c5.29,0,9.41,1.33,12.35,4c2.94,2.67,4.42,6.39,4.42,11.18c0,4.78-1.47,8.51-4.42,11.18\n c-2.94,2.67-7.06,4-12.35,4H34.59v18.29H23.83V125.68z M44.81,147.9c5.38,0,8.07-2.32,8.07-6.97c0-2.39-0.67-4.16-2-5.31\n c-1.33-1.15-3.36-1.73-6.07-1.73H34.59v14.01H44.81z\"\n />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M69.92,125.68h18.91c5.29,0,9.84,0.97,13.66,2.9c3.82,1.93,6.74,4.72,8.76,8.35\n c2.02,3.63,3.04,7.98,3.04,13.04c0,5.06-1,9.42-3,13.08c-2,3.66-4.91,6.45-8.73,8.38c-3.82,1.93-8.4,2.9-13.73,2.9H69.92V125.68z\n M88.07,165.63c10.35,0,15.52-5.22,15.52-15.66c0-10.4-5.17-15.59-15.52-15.59h-7.38v31.26H88.07z\"\n />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M122.57,125.68h32.84v8.49h-22.22v11.18h20.84v8.49h-20.84v20.49h-10.63V125.68z\"\n />\n </g>\n </svg>\n <span class=\"product-documents__info\"\n >Technical Datasheet\n </span>\n </a>\n </div>\n <div class=\"product-documents\">\n <a\n href=\"/content/documents/manuals/cut-and-run-kit-manual.pdf\"\n target=\"_new\"\n class=\"product-documents__link\"\n >\n <svg\n version=\"1.1\"\n id=\"Layer_1\"\n xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\"\n x=\"0px\"\n y=\"0px\"\n viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\"\n xml:space=\"preserve\"\n class=\"product-documents__icon\"\n alt=\"16-0002 Datasheet\"\n >\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M191.92,68.77l-47.69-47.69c-1.33-1.33-3.12-2.08-5.01-2.08H45.09C41.17,19,38,22.17,38,26.09v184.36\n c0,3.92,3.17,7.09,7.09,7.09h141.82c3.92,0,7.09-3.17,7.09-7.09V73.8C194,71.92,193.25,70.1,191.92,68.77z M177.65,77.06h-41.7\n v-41.7L177.65,77.06z M178.05,201.59H53.95V34.95h66.92v47.86c0,5.14,4.17,9.31,9.31,9.31h47.86V201.59z\"\n />\n </g>\n <rect\n x=\"20\"\n y=\"112\"\n class=\"product-documents__svg-background\"\n width=\"146\"\n height=\"76\"\n />\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M23.83,125.68h22.36c5.29,0,9.41,1.33,12.35,4c2.94,2.67,4.42,6.39,4.42,11.18c0,4.78-1.47,8.51-4.42,11.18\n c-2.94,2.67-7.06,4-12.35,4H34.59v18.29H23.83V125.68z M44.81,147.9c5.38,0,8.07-2.32,8.07-6.97c0-2.39-0.67-4.16-2-5.31\n c-1.33-1.15-3.36-1.73-6.07-1.73H34.59v14.01H44.81z\"\n />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M69.92,125.68h18.91c5.29,0,9.84,0.97,13.66,2.9c3.82,1.93,6.74,4.72,8.76,8.35\n c2.02,3.63,3.04,7.98,3.04,13.04c0,5.06-1,9.42-3,13.08c-2,3.66-4.91,6.45-8.73,8.38c-3.82,1.93-8.4,2.9-13.73,2.9H69.92V125.68z\n M88.07,165.63c10.35,0,15.52-5.22,15.52-15.66c0-10.4-5.17-15.59-15.52-15.59h-7.38v31.26H88.07z\"\n />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M122.57,125.68h32.84v8.49h-22.22v11.18h20.84v8.49h-20.84v20.49h-10.63V125.68z\"\n />\n </g>\n </svg>\n <span class=\"product-documents__info\"\n >CUT&RUN Kit Manual</span\n >\n </a>\n </div>\n <div class=\"product-documents\">\n <a\n href=\"/content/documents/manuals/cut-and-run-kit-quick-card.pdf\"\n target=\"_new\"\n class=\"product-documents__link\"\n >\n <svg\n version=\"1.1\"\n id=\"Layer_1\"\n xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\"\n x=\"0px\"\n y=\"0px\"\n viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\"\n xml:space=\"preserve\"\n class=\"product-documents__icon\"\n alt=\"16-0002 Datasheet\"\n >\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M191.92,68.77l-47.69-47.69c-1.33-1.33-3.12-2.08-5.01-2.08H45.09C41.17,19,38,22.17,38,26.09v184.36\n c0,3.92,3.17,7.09,7.09,7.09h141.82c3.92,0,7.09-3.17,7.09-7.09V73.8C194,71.92,193.25,70.1,191.92,68.77z M177.65,77.06h-41.7\n v-41.7L177.65,77.06z M178.05,201.59H53.95V34.95h66.92v47.86c0,5.14,4.17,9.31,9.31,9.31h47.86V201.59z\"\n />\n </g>\n <rect\n x=\"20\"\n y=\"112\"\n class=\"product-documents__svg-background\"\n width=\"146\"\n height=\"76\"\n />\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M23.83,125.68h22.36c5.29,0,9.41,1.33,12.35,4c2.94,2.67,4.42,6.39,4.42,11.18c0,4.78-1.47,8.51-4.42,11.18\n c-2.94,2.67-7.06,4-12.35,4H34.59v18.29H23.83V125.68z M44.81,147.9c5.38,0,8.07-2.32,8.07-6.97c0-2.39-0.67-4.16-2-5.31\n c-1.33-1.15-3.36-1.73-6.07-1.73H34.59v14.01H44.81z\"\n />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M69.92,125.68h18.91c5.29,0,9.84,0.97,13.66,2.9c3.82,1.93,6.74,4.72,8.76,8.35\n c2.02,3.63,3.04,7.98,3.04,13.04c0,5.06-1,9.42-3,13.08c-2,3.66-4.91,6.45-8.73,8.38c-3.82,1.93-8.4,2.9-13.73,2.9H69.92V125.68z\n M88.07,165.63c10.35,0,15.52-5.22,15.52-15.66c0-10.4-5.17-15.59-15.52-15.59h-7.38v31.26H88.07z\"\n />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M122.57,125.68h32.84v8.49h-22.22v11.18h20.84v8.49h-20.84v20.49h-10.63V125.68z\"\n />\n </g>\n </svg>\n <span class=\"product-documents__info\"\n >CUT&RUN Kit Quick Card</span\n >\n </a>\n </div>\n <div class=\"product-documents\">\n <a\n href=\"/content/documents/SNAP-CUTANA_K-MetStat_Panel_ShellScript.sh\"\n target=\"_blank\"\n class=\"product-documents__link\"\n >\n <svg\n version=\"1.1\"\n id=\"Layer_1\"\n xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\"\n x=\"0px\"\n y=\"0px\"\n viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\"\n xml:space=\"preserve\"\n class=\"product-documents__icon\"\n >\n <g>\n <path\n class=\"product-documents__svg-txt\"\n d=\"M199.12,66.39l-51.16-51.16c-1.43-1.43-3.35-2.23-5.37-2.23H41.61c-4.2,0-7.61,3.41-7.61,7.61v197.79\n c0,4.2,3.41,7.61,7.61,7.61h152.14c4.2,0,7.61-3.41,7.61-7.61V71.79C201.36,69.77,200.55,67.82,199.12,66.39L199.12,66.39z\n M183.81,75.28h-44.74V30.54L183.81,75.28z M184.24,208.88H51.12V30.12h71.79v51.35c0,5.51,4.47,9.98,9.98,9.98h51.35V208.88z\n M115.78,144.7H72.04c-1.05,0-1.9,0.86-1.9,1.9v11.41c0,1.05,0.86,1.9,1.9,1.9h43.74c1.05,0,1.9-0.86,1.9-1.9V146.6\n C117.68,145.55,116.82,144.7,115.78,144.7z M70.13,114.27v11.41c0,1.05,0.86,1.9,1.9,1.9h91.29c1.05,0,1.9-0.86,1.9-1.9v-11.41\n c0-1.05-0.86-1.9-1.9-1.9H72.04C70.99,112.37,70.13,113.22,70.13,114.27z\"\n />\n </g>\n </svg>\n <span class=\"product-documents__info\">\n SNAP-CUTANA™ K-MetStat Panel Shell Script</span\n >\n </a>\n </div>\n <div class=\"product-documents\">\n <a\n href=\"/content/documents/SNAP-CUTANA_K-MetStat_Panel_Analysis.xlsx\"\n target=\"_blank\"\n class=\"product-documents__link\"\n >\n <svg\n version=\"1.1\"\n id=\"Layer_1\"\n xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\"\n x=\"0px\"\n y=\"0px\"\n viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\"\n xml:space=\"preserve\"\n class=\"product-documents__icon\"\n >\n <g>\n <path\n class=\"product-documents__svg-excel\"\n d=\"M193.17,69.88l-46.83-46.83c-1.31-1.31-3.07-2.05-4.92-2.05H48.96C45.11,21,42,24.11,42,27.96v181.07\n c0,3.85,3.11,6.96,6.96,6.96h139.29c3.85,0,6.96-3.11,6.96-6.96V74.82C195.21,72.97,194.47,71.19,193.17,69.88L193.17,69.88z\n M179.15,78.02h-40.96V37.06L179.15,78.02z M179.54,200.33H57.67V36.67h65.73v47.01c0,5.05,4.09,9.14,9.14,9.14h47.01V200.33z\n M119.06,133.32l-13.45-22.29c-0.48-0.78-2.24-1.24-2.24-1.24h-8.33c-0.5,0-0.98,0.13-1.39,0.41c-1.21,0.76-1.58,2.36-0.8,3.6\n l17.85,28.28l-18.08,28.8c-0.76,1.22-0.39,2.83,0.84,3.6c0.41,0.26,0.89,0.39,1.37,0.39h7.48c0,0,1.74-0.26,2.22-1.02l13.65-22.09\n l13.56,22.07c0.48,0.78,2.22,1.26,2.22,1.26h8.18c0.5,0,0.98-0.15,1.42-0.42c1.22-0.79,1.57-2.4,0.79-3.63l-18.35-28.48\n l18.63-28.94c0.78-1.23,0.42-2.85-0.81-3.63c-0.42-0.27-0.9-0.41-1.4-0.41h-7.79c0,0-1.76,0.7-2.24,1.48L119.06,133.32z\"\n />\n </g>\n </svg>\n <span class=\"product-documents__info\"\n >SNAP-CUTANA™ K-MetStat Panel Analysis</span\n >\n </a>\n </div>\n </div>\n </div>\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <!-- <h3 style=\"color: #4698cb; margin-bottom: 0\">Current Lot</h3> -->\n <h3 style=\"margin-top: 1rem; padding-top: 0\" class=\"sub-title1\">\n Kit Version 3\n </h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\"\n >\n <div class=\"product-documents\">\n <a\n href=\"/content/documents/tds/14-1048-3.5.pdf\"\n target=\"_new\"\n class=\"product-documents__link\"\n >\n <svg\n version=\"1.1\"\n id=\"Layer_1\"\n xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\"\n x=\"0px\"\n y=\"0px\"\n viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\"\n xml:space=\"preserve\"\n class=\"product-documents__icon\"\n alt=\"16-0002 Datasheet\"\n >\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M191.92,68.77l-47.69-47.69c-1.33-1.33-3.12-2.08-5.01-2.08H45.09C41.17,19,38,22.17,38,26.09v184.36\n c0,3.92,3.17,7.09,7.09,7.09h141.82c3.92,0,7.09-3.17,7.09-7.09V73.8C194,71.92,193.25,70.1,191.92,68.77z M177.65,77.06h-41.7\n v-41.7L177.65,77.06z M178.05,201.59H53.95V34.95h66.92v47.86c0,5.14,4.17,9.31,9.31,9.31h47.86V201.59z\"\n />\n </g>\n <rect\n x=\"20\"\n y=\"112\"\n class=\"product-documents__svg-background\"\n width=\"146\"\n height=\"76\"\n />\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M23.83,125.68h22.36c5.29,0,9.41,1.33,12.35,4c2.94,2.67,4.42,6.39,4.42,11.18c0,4.78-1.47,8.51-4.42,11.18\n c-2.94,2.67-7.06,4-12.35,4H34.59v18.29H23.83V125.68z M44.81,147.9c5.38,0,8.07-2.32,8.07-6.97c0-2.39-0.67-4.16-2-5.31\n c-1.33-1.15-3.36-1.73-6.07-1.73H34.59v14.01H44.81z\"\n />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M69.92,125.68h18.91c5.29,0,9.84,0.97,13.66,2.9c3.82,1.93,6.74,4.72,8.76,8.35\n c2.02,3.63,3.04,7.98,3.04,13.04c0,5.06-1,9.42-3,13.08c-2,3.66-4.91,6.45-8.73,8.38c-3.82,1.93-8.4,2.9-13.73,2.9H69.92V125.68z\n M88.07,165.63c10.35,0,15.52-5.22,15.52-15.66c0-10.4-5.17-15.59-15.52-15.59h-7.38v31.26H88.07z\"\n />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M122.57,125.68h32.84v8.49h-22.22v11.18h20.84v8.49h-20.84v20.49h-10.63V125.68z\"\n />\n </g>\n </svg>\n <span class=\"product-documents__info\"\n >Technical Datasheet\n </span>\n </a>\n </div>\n <div class=\"product-documents\">\n <a\n href=\"/content/documents/manuals/cut-and-run-kit-manual_3.5.pdf\"\n target=\"_new\"\n class=\"product-documents__link\"\n >\n <svg\n version=\"1.1\"\n id=\"Layer_1\"\n xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\"\n x=\"0px\"\n y=\"0px\"\n viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\"\n xml:space=\"preserve\"\n class=\"product-documents__icon\"\n alt=\"16-0002 Datasheet\"\n >\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M191.92,68.77l-47.69-47.69c-1.33-1.33-3.12-2.08-5.01-2.08H45.09C41.17,19,38,22.17,38,26.09v184.36\n c0,3.92,3.17,7.09,7.09,7.09h141.82c3.92,0,7.09-3.17,7.09-7.09V73.8C194,71.92,193.25,70.1,191.92,68.77z M177.65,77.06h-41.7\n v-41.7L177.65,77.06z M178.05,201.59H53.95V34.95h66.92v47.86c0,5.14,4.17,9.31,9.31,9.31h47.86V201.59z\"\n />\n </g>\n <rect\n x=\"20\"\n y=\"112\"\n class=\"product-documents__svg-background\"\n width=\"146\"\n height=\"76\"\n />\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M23.83,125.68h22.36c5.29,0,9.41,1.33,12.35,4c2.94,2.67,4.42,6.39,4.42,11.18c0,4.78-1.47,8.51-4.42,11.18\n c-2.94,2.67-7.06,4-12.35,4H34.59v18.29H23.83V125.68z M44.81,147.9c5.38,0,8.07-2.32,8.07-6.97c0-2.39-0.67-4.16-2-5.31\n c-1.33-1.15-3.36-1.73-6.07-1.73H34.59v14.01H44.81z\"\n />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M69.92,125.68h18.91c5.29,0,9.84,0.97,13.66,2.9c3.82,1.93,6.74,4.72,8.76,8.35\n c2.02,3.63,3.04,7.98,3.04,13.04c0,5.06-1,9.42-3,13.08c-2,3.66-4.91,6.45-8.73,8.38c-3.82,1.93-8.4,2.9-13.73,2.9H69.92V125.68z\n M88.07,165.63c10.35,0,15.52-5.22,15.52-15.66c0-10.4-5.17-15.59-15.52-15.59h-7.38v31.26H88.07z\"\n />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M122.57,125.68h32.84v8.49h-22.22v11.18h20.84v8.49h-20.84v20.49h-10.63V125.68z\"\n />\n </g>\n </svg>\n <span class=\"product-documents__info\"\n >CUT&RUN Kit Manual</span\n >\n </a>\n </div>\n <div class=\"product-documents\">\n <a\n href=\"/content/documents/manuals/cut-and-run-kit-quick-card_3.5.pdf\"\n target=\"_new\"\n class=\"product-documents__link\"\n >\n <svg\n version=\"1.1\"\n id=\"Layer_1\"\n xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\"\n x=\"0px\"\n y=\"0px\"\n viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\"\n xml:space=\"preserve\"\n class=\"product-documents__icon\"\n alt=\"16-0002 Datasheet\"\n >\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M191.92,68.77l-47.69-47.69c-1.33-1.33-3.12-2.08-5.01-2.08H45.09C41.17,19,38,22.17,38,26.09v184.36\n c0,3.92,3.17,7.09,7.09,7.09h141.82c3.92,0,7.09-3.17,7.09-7.09V73.8C194,71.92,193.25,70.1,191.92,68.77z M177.65,77.06h-41.7\n v-41.7L177.65,77.06z M178.05,201.59H53.95V34.95h66.92v47.86c0,5.14,4.17,9.31,9.31,9.31h47.86V201.59z\"\n />\n </g>\n <rect\n x=\"20\"\n y=\"112\"\n class=\"product-documents__svg-background\"\n width=\"146\"\n height=\"76\"\n />\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M23.83,125.68h22.36c5.29,0,9.41,1.33,12.35,4c2.94,2.67,4.42,6.39,4.42,11.18c0,4.78-1.47,8.51-4.42,11.18\n c-2.94,2.67-7.06,4-12.35,4H34.59v18.29H23.83V125.68z M44.81,147.9c5.38,0,8.07-2.32,8.07-6.97c0-2.39-0.67-4.16-2-5.31\n c-1.33-1.15-3.36-1.73-6.07-1.73H34.59v14.01H44.81z\"\n />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M69.92,125.68h18.91c5.29,0,9.84,0.97,13.66,2.9c3.82,1.93,6.74,4.72,8.76,8.35\n c2.02,3.63,3.04,7.98,3.04,13.04c0,5.06-1,9.42-3,13.08c-2,3.66-4.91,6.45-8.73,8.38c-3.82,1.93-8.4,2.9-13.73,2.9H69.92V125.68z\n M88.07,165.63c10.35,0,15.52-5.22,15.52-15.66c0-10.4-5.17-15.59-15.52-15.59h-7.38v31.26H88.07z\"\n />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M122.57,125.68h32.84v8.49h-22.22v11.18h20.84v8.49h-20.84v20.49h-10.63V125.68z\"\n />\n </g>\n </svg>\n <span class=\"product-documents__info\"\n >CUT&RUN Kit Quick Card</span\n >\n </a>\n </div>\n <div class=\"product-documents\">\n <a\n href=\"/content/documents/SNAP-CUTANA_K-MetStat_Panel_ShellScript_2.sh\"\n target=\"_blank\"\n class=\"product-documents__link\"\n >\n <svg\n version=\"1.1\"\n id=\"Layer_1\"\n xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\"\n x=\"0px\"\n y=\"0px\"\n viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\"\n xml:space=\"preserve\"\n class=\"product-documents__icon\"\n >\n <g>\n <path\n class=\"product-documents__svg-txt\"\n d=\"M199.12,66.39l-51.16-51.16c-1.43-1.43-3.35-2.23-5.37-2.23H41.61c-4.2,0-7.61,3.41-7.61,7.61v197.79\n c0,4.2,3.41,7.61,7.61,7.61h152.14c4.2,0,7.61-3.41,7.61-7.61V71.79C201.36,69.77,200.55,67.82,199.12,66.39L199.12,66.39z\n M183.81,75.28h-44.74V30.54L183.81,75.28z M184.24,208.88H51.12V30.12h71.79v51.35c0,5.51,4.47,9.98,9.98,9.98h51.35V208.88z\n M115.78,144.7H72.04c-1.05,0-1.9,0.86-1.9,1.9v11.41c0,1.05,0.86,1.9,1.9,1.9h43.74c1.05,0,1.9-0.86,1.9-1.9V146.6\n C117.68,145.55,116.82,144.7,115.78,144.7z M70.13,114.27v11.41c0,1.05,0.86,1.9,1.9,1.9h91.29c1.05,0,1.9-0.86,1.9-1.9v-11.41\n c0-1.05-0.86-1.9-1.9-1.9H72.04C70.99,112.37,70.13,113.22,70.13,114.27z\"\n />\n </g>\n </svg>\n <span class=\"product-documents__info\">\n SNAP-CUTANA™ K-MetStat Panel Shell Script</span\n >\n </a>\n </div>\n <div class=\"product-documents\">\n <a\n href=\"/content/documents/SNAP-CUTANA_K-MetStat_Panel_Analysis_3.xlsx\"\n target=\"_blank\"\n class=\"product-documents__link\"\n >\n <svg\n version=\"1.1\"\n id=\"Layer_1\"\n xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\"\n x=\"0px\"\n y=\"0px\"\n viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\"\n xml:space=\"preserve\"\n class=\"product-documents__icon\"\n >\n <g>\n <path\n class=\"product-documents__svg-excel\"\n d=\"M193.17,69.88l-46.83-46.83c-1.31-1.31-3.07-2.05-4.92-2.05H48.96C45.11,21,42,24.11,42,27.96v181.07\n c0,3.85,3.11,6.96,6.96,6.96h139.29c3.85,0,6.96-3.11,6.96-6.96V74.82C195.21,72.97,194.47,71.19,193.17,69.88L193.17,69.88z\n M179.15,78.02h-40.96V37.06L179.15,78.02z M179.54,200.33H57.67V36.67h65.73v47.01c0,5.05,4.09,9.14,9.14,9.14h47.01V200.33z\n M119.06,133.32l-13.45-22.29c-0.48-0.78-2.24-1.24-2.24-1.24h-8.33c-0.5,0-0.98,0.13-1.39,0.41c-1.21,0.76-1.58,2.36-0.8,3.6\n l17.85,28.28l-18.08,28.8c-0.76,1.22-0.39,2.83,0.84,3.6c0.41,0.26,0.89,0.39,1.37,0.39h7.48c0,0,1.74-0.26,2.22-1.02l13.65-22.09\n l13.56,22.07c0.48,0.78,2.22,1.26,2.22,1.26h8.18c0.5,0,0.98-0.15,1.42-0.42c1.22-0.79,1.57-2.4,0.79-3.63l-18.35-28.48\n l18.63-28.94c0.78-1.23,0.42-2.85-0.81-3.63c-0.42-0.27-0.9-0.41-1.4-0.41h-7.79c0,0-1.76,0.7-2.24,1.48L119.06,133.32z\"\n />\n </g>\n </svg>\n <span class=\"product-documents__info\"\n >SNAP-CUTANA™ K-MetStat Panel Analysis</span\n >\n </a>\n </div>\n </div>\n </div>\n </div>\n <!-- <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Kit Version 2</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\">\n <div class=\"product-documents\">\n <a\n href=\"/content/documents/tds/14-1048-21130003-01.pdf\"\n target=\"_new\"\n class=\"product-documents__link\">\n <svg\n version=\"1.1\"\n id=\"Layer_1\"\n xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\"\n x=\"0px\"\n y=\"0px\"\n viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\"\n xml:space=\"preserve\"\n class=\"product-documents__icon\"\n alt=\"16-0002 Datasheet\">\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M191.92,68.77l-47.69-47.69c-1.33-1.33-3.12-2.08-5.01-2.08H45.09C41.17,19,38,22.17,38,26.09v184.36\n c0,3.92,3.17,7.09,7.09,7.09h141.82c3.92,0,7.09-3.17,7.09-7.09V73.8C194,71.92,193.25,70.1,191.92,68.77z M177.65,77.06h-41.7\n v-41.7L177.65,77.06z M178.05,201.59H53.95V34.95h66.92v47.86c0,5.14,4.17,9.31,9.31,9.31h47.86V201.59z\" />\n </g>\n <rect\n x=\"20\"\n y=\"112\"\n class=\"product-documents__svg-background\"\n width=\"146\"\n height=\"76\" />\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M23.83,125.68h22.36c5.29,0,9.41,1.33,12.35,4c2.94,2.67,4.42,6.39,4.42,11.18c0,4.78-1.47,8.51-4.42,11.18\n c-2.94,2.67-7.06,4-12.35,4H34.59v18.29H23.83V125.68z M44.81,147.9c5.38,0,8.07-2.32,8.07-6.97c0-2.39-0.67-4.16-2-5.31\n c-1.33-1.15-3.36-1.73-6.07-1.73H34.59v14.01H44.81z\" />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M69.92,125.68h18.91c5.29,0,9.84,0.97,13.66,2.9c3.82,1.93,6.74,4.72,8.76,8.35\n c2.02,3.63,3.04,7.98,3.04,13.04c0,5.06-1,9.42-3,13.08c-2,3.66-4.91,6.45-8.73,8.38c-3.82,1.93-8.4,2.9-13.73,2.9H69.92V125.68z\n M88.07,165.63c10.35,0,15.52-5.22,15.52-15.66c0-10.4-5.17-15.59-15.52-15.59h-7.38v31.26H88.07z\" />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M122.57,125.68h32.84v8.49h-22.22v11.18h20.84v8.49h-20.84v20.49h-10.63V125.68z\" />\n </g>\n </svg>\n <span class=\"product-documents__info\">Technical Datasheet</span>\n </a>\n </div>\n\n <div class=\"product-documents\">\n <a\n href=\"/content/documents/manuals/cut-and-run-kit_2.1.pdf\"\n target=\"_new\"\n class=\"product-documents__link\">\n <svg\n version=\"1.1\"\n id=\"Layer_1\"\n xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\"\n x=\"0px\"\n y=\"0px\"\n viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\"\n xml:space=\"preserve\"\n class=\"product-documents__icon\"\n alt=\"16-0002 Datasheet\">\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M191.92,68.77l-47.69-47.69c-1.33-1.33-3.12-2.08-5.01-2.08H45.09C41.17,19,38,22.17,38,26.09v184.36\n c0,3.92,3.17,7.09,7.09,7.09h141.82c3.92,0,7.09-3.17,7.09-7.09V73.8C194,71.92,193.25,70.1,191.92,68.77z M177.65,77.06h-41.7\n v-41.7L177.65,77.06z M178.05,201.59H53.95V34.95h66.92v47.86c0,5.14,4.17,9.31,9.31,9.31h47.86V201.59z\" />\n </g>\n <rect\n x=\"20\"\n y=\"112\"\n class=\"product-documents__svg-background\"\n width=\"146\"\n height=\"76\" />\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M23.83,125.68h22.36c5.29,0,9.41,1.33,12.35,4c2.94,2.67,4.42,6.39,4.42,11.18c0,4.78-1.47,8.51-4.42,11.18\n c-2.94,2.67-7.06,4-12.35,4H34.59v18.29H23.83V125.68z M44.81,147.9c5.38,0,8.07-2.32,8.07-6.97c0-2.39-0.67-4.16-2-5.31\n c-1.33-1.15-3.36-1.73-6.07-1.73H34.59v14.01H44.81z\" />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M69.92,125.68h18.91c5.29,0,9.84,0.97,13.66,2.9c3.82,1.93,6.74,4.72,8.76,8.35\n c2.02,3.63,3.04,7.98,3.04,13.04c0,5.06-1,9.42-3,13.08c-2,3.66-4.91,6.45-8.73,8.38c-3.82,1.93-8.4,2.9-13.73,2.9H69.92V125.68z\n M88.07,165.63c10.35,0,15.52-5.22,15.52-15.66c0-10.4-5.17-15.59-15.52-15.59h-7.38v31.26H88.07z\" />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M122.57,125.68h32.84v8.49h-22.22v11.18h20.84v8.49h-20.84v20.49h-10.63V125.68z\" />\n </g>\n </svg>\n <span class=\"product-documents__info\">CUT&RUN Kit Manual</span>\n </a>\n </div>\n <div class=\"product-documents\">\n <a\n href=\"/content/documents/manuals/cut-and-run-kit-quick-card_2.0.pdf\"\n target=\"_new\"\n class=\"product-documents__link\">\n <svg\n version=\"1.1\"\n id=\"Layer_1\"\n xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\"\n x=\"0px\"\n y=\"0px\"\n viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\"\n xml:space=\"preserve\"\n class=\"product-documents__icon\"\n alt=\"16-0002 Datasheet\">\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M191.92,68.77l-47.69-47.69c-1.33-1.33-3.12-2.08-5.01-2.08H45.09C41.17,19,38,22.17,38,26.09v184.36\n c0,3.92,3.17,7.09,7.09,7.09h141.82c3.92,0,7.09-3.17,7.09-7.09V73.8C194,71.92,193.25,70.1,191.92,68.77z M177.65,77.06h-41.7\n v-41.7L177.65,77.06z M178.05,201.59H53.95V34.95h66.92v47.86c0,5.14,4.17,9.31,9.31,9.31h47.86V201.59z\" />\n </g>\n <rect\n x=\"20\"\n y=\"112\"\n class=\"product-documents__svg-background\"\n width=\"146\"\n height=\"76\" />\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M23.83,125.68h22.36c5.29,0,9.41,1.33,12.35,4c2.94,2.67,4.42,6.39,4.42,11.18c0,4.78-1.47,8.51-4.42,11.18\n c-2.94,2.67-7.06,4-12.35,4H34.59v18.29H23.83V125.68z M44.81,147.9c5.38,0,8.07-2.32,8.07-6.97c0-2.39-0.67-4.16-2-5.31\n c-1.33-1.15-3.36-1.73-6.07-1.73H34.59v14.01H44.81z\" />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M69.92,125.68h18.91c5.29,0,9.84,0.97,13.66,2.9c3.82,1.93,6.74,4.72,8.76,8.35\n c2.02,3.63,3.04,7.98,3.04,13.04c0,5.06-1,9.42-3,13.08c-2,3.66-4.91,6.45-8.73,8.38c-3.82,1.93-8.4,2.9-13.73,2.9H69.92V125.68z\n M88.07,165.63c10.35,0,15.52-5.22,15.52-15.66c0-10.4-5.17-15.59-15.52-15.59h-7.38v31.26H88.07z\" />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M122.57,125.68h32.84v8.49h-22.22v11.18h20.84v8.49h-20.84v20.49h-10.63V125.68z\" />\n </g>\n </svg>\n <span class=\"product-documents__info\"\n >CUT&RUN Kit Quick Card</span\n >\n </a>\n </div>\n <div class=\"product-documents\">\n <a\n href=\"/content/documents/14-1048-H3K4MetStat-Shell-Script_2.sh\"\n target=\"_blank\"\n class=\"product-documents__link\">\n <svg\n version=\"1.1\"\n id=\"Layer_1\"\n xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\"\n x=\"0px\"\n y=\"0px\"\n viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\"\n xml:space=\"preserve\"\n class=\"product-documents__icon\">\n <g>\n <path\n class=\"product-documents__svg-txt\"\n d=\"M199.12,66.39l-51.16-51.16c-1.43-1.43-3.35-2.23-5.37-2.23H41.61c-4.2,0-7.61,3.41-7.61,7.61v197.79\n c0,4.2,3.41,7.61,7.61,7.61h152.14c4.2,0,7.61-3.41,7.61-7.61V71.79C201.36,69.77,200.55,67.82,199.12,66.39L199.12,66.39z\n M183.81,75.28h-44.74V30.54L183.81,75.28z M184.24,208.88H51.12V30.12h71.79v51.35c0,5.51,4.47,9.98,9.98,9.98h51.35V208.88z\n M115.78,144.7H72.04c-1.05,0-1.9,0.86-1.9,1.9v11.41c0,1.05,0.86,1.9,1.9,1.9h43.74c1.05,0,1.9-0.86,1.9-1.9V146.6\n C117.68,145.55,116.82,144.7,115.78,144.7z M70.13,114.27v11.41c0,1.05,0.86,1.9,1.9,1.9h91.29c1.05,0,1.9-0.86,1.9-1.9v-11.41\n c0-1.05-0.86-1.9-1.9-1.9H72.04C70.99,112.37,70.13,113.22,70.13,114.27z\" />\n </g>\n </svg>\n <span class=\"product-documents__info\">\n H3K4 MetStat Spike-in Controls Shell Script</span\n >\n </a>\n </div>\n <div class=\"product-documents\">\n <a\n href=\"/content/documents/H3K4-MetStat-Spike-in-Controls-Analysis_2.xlsx\"\n target=\"_blank\"\n class=\"product-documents__link\">\n <svg\n version=\"1.1\"\n id=\"Layer_1\"\n xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\"\n x=\"0px\"\n y=\"0px\"\n viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\"\n xml:space=\"preserve\"\n class=\"product-documents__icon\">\n <g>\n <path\n class=\"product-documents__svg-excel\"\n d=\"M193.17,69.88l-46.83-46.83c-1.31-1.31-3.07-2.05-4.92-2.05H48.96C45.11,21,42,24.11,42,27.96v181.07\n c0,3.85,3.11,6.96,6.96,6.96h139.29c3.85,0,6.96-3.11,6.96-6.96V74.82C195.21,72.97,194.47,71.19,193.17,69.88L193.17,69.88z\n M179.15,78.02h-40.96V37.06L179.15,78.02z M179.54,200.33H57.67V36.67h65.73v47.01c0,5.05,4.09,9.14,9.14,9.14h47.01V200.33z\n M119.06,133.32l-13.45-22.29c-0.48-0.78-2.24-1.24-2.24-1.24h-8.33c-0.5,0-0.98,0.13-1.39,0.41c-1.21,0.76-1.58,2.36-0.8,3.6\n l17.85,28.28l-18.08,28.8c-0.76,1.22-0.39,2.83,0.84,3.6c0.41,0.26,0.89,0.39,1.37,0.39h7.48c0,0,1.74-0.26,2.22-1.02l13.65-22.09\n l13.56,22.07c0.48,0.78,2.22,1.26,2.22,1.26h8.18c0.5,0,0.98-0.15,1.42-0.42c1.22-0.79,1.57-2.4,0.79-3.63l-18.35-28.48\n l18.63-28.94c0.78-1.23,0.42-2.85-0.81-3.63c-0.42-0.27-0.9-0.41-1.4-0.41h-7.79c0,0-1.76,0.7-2.24,1.48L119.06,133.32z\" />\n </g>\n </svg>\n <span class=\"product-documents__info\"\n >H3K4 MetStat Spike-in Controls Analysis</span\n >\n </a>\n </div>\n </div>\n </div>\n </div> -->\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel current\">\n <h3 id=\"bioz\" class=\"sub-title1\">Product References</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description-specific\"\n >\n <object\n id=\"wobj-3835-14-1048-q\"\n type=\"text/html\"\n data=\"https://www.bioz.com/v_widget_6_0/3835/14-1048/?ex=1\"\n style=\"width: 100%; height: 193px\"\n ></object>\n <div id=\"bioz-w-pb-3835-14-1048-q-div\" style=\"width: 100%\">\n <a\n id=\"bioz-w-pb-3835-14-1048-q\"\n style=\"\n font-size: 12px;\n text-decoration: none;\n color: #4698cb;\n \"\n href=\"https://www.bioz.com/\"\n target=\"_blank\"\n ><img\n src=\"https://cdn.bioz.com/assets/favicon.png\"\n style=\"\n width: 11px;\n height: 11px;\n vertical-align: baseline;\n padding-bottom: 0px;\n margin-left: 0px;\n margin-bottom: 0px;\n float: none;\n \"\n />\n Powered by Bioz</a\n >\n <a\n style=\"\n font-size: 12px;\n text-decoration: none;\n float: right;\n color: transparent;\n \"\n href=\"https://www.bioz.com/result/14-1048/product/EpiCypher/?cn=14-1048\"\n target=\"_blank\"\n >\n See more details on Bioz</a\n >\n </div>\n </div>\n </div>\n </div>\n </div>\n</div>\n\n<script>\n $(document).ready(function () {\n var widget_micro_obj = new v_widget_obj(\"s\", [1]);\n widget_micro_obj.request_catalog_number_widget_data_internal(\n \"14-1048\",\n \"14-1048\"\n );\n });\n</script>\n\n<style>\n .form-field-title .required-text {\n display: none !important;\n }\n\n .form-label {\n padding-left: 2rem;\n }\n\n .form-label-text {\n margin: 0;\n margin-left: 0 !important;\n }\n\n /* //////////// */\n\n /* BIOZ */\n td table {\n margin: 0;\n padding: 6px !important;\n }\n\n span.bioz-w-parent-hover {\n font-size: 15px;\n }\n\n td .bioz-w-tooltipx {\n padding-top: 0 !important;\n }\n /* .bioz-w-parent-hover:hover {\n text-decoration: underline white 2px !important;\n } */\n\n /* /////////// */\n\n .form-field-title .required-text {\n display: none !important;\n }\n .form-label {\n padding-left: 2rem;\n }\n .form-label-text {\n margin: 0;\n margin-left: 0 !important;\n }\n\n .epicypher-table table {\n margin-right: auto;\n margin-left: auto;\n width: 100%;\n border-top: 1px solid lightgray;\n }\n\n .epicypher-table th {\n color: #3b3a48;\n }\n .epicypher-table td {\n color: #3b3a48;\n font-size: 0.9rem;\n }\n .epicypher-table td a {\n text-decoration: underline;\n }\n .epicypher-table td,\n th {\n border: 0.5px solid lightgray;\n text-align: left;\n padding: 8px;\n /* white-space: nowrap; */\n }\n .epicypher-table tr:nth-child(even) {\n background-color: #f9f9f9;\n }\n .epicypher-table td:first-child {\n border-left: 1px solid lightgray;\n }\n\n /* Tablet */\n\n @media only screen and (max-width: 1024px) {\n /* Force table to not be like tables anymore */\n .epicypher-table th {\n display: none !important;\n }\n\n /* BIOZ */\n td.bioz-w-tooltipx::before {\n display: none;\n }\n\n /* Hide table headers (but not display: none;, for accessibility) */\n .epicypher-table thead tr {\n position: absolute;\n top: -9999px;\n left: -9999px;\n }\n\n .epicypher-table tr {\n border: 1px solid #ccc;\n }\n\n .epicypher-table td {\n /* Behave like a \"row\" */\n /* border: none; */\n border-bottom: 1px solid #eee;\n position: relative;\n padding-left: 30%;\n white-space: inherit;\n }\n\n .epicypher-table td:before {\n /* Now like a table header */\n position: absolute;\n /* Top/left values mimic padding */\n top: 6px;\n left: 6px;\n width: 45%;\n padding-right: 10px;\n white-space: nowrap;\n font-weight: 700;\n }\n\n /*\n Label the data\n */\n .epicypher-table td:nth-of-type(1):before {\n content: \"Item\";\n }\n td:nth-of-type(2):before {\n content: \"Cat. No.\";\n }\n }\n\n @media screen and (min-width: 768px) {\n .products-featured .container,\n .products-featured .product-tabs,\n .products-related .container,\n .products-related .product-tabs {\n padding-left: 30px !important;\n padding-right: 50px;\n max-width: 1170px !important;\n }\n }\n\n @media screen and (max-width: 767px) {\n .custom-title-description {\n font-size: 19px !important;\n }\n\n .section-title {\n font-size: 19px !important;\n padding-left: 1rem;\n }\n }\n</style>","tags":[],"warranty":"","price":{"without_tax":{"formatted":"$945.00","value":945,"currency":"USD"},"tax_label":"Sales Tax"},"detail_messages":"","availability":"","page_title":"CUTANA™ ChIC/CUT&RUN Kit","cart_url":"https://www.epicypher.com/cart.php","max_purchase_quantity":0,"mpn":"","upc":null,"options":[],"related_products":[{"id":694,"sku":null,"name":"CUTANA™ pAG-MNase for ChIC/CUT&RUN Workflows","url":"https://www.epicypher.com/products/epigenetics-reagents-and-assays/cutana-pag-mnase-for-chic-cut-and-run-workflows","availability":"","rating":null,"brand":{"name":null},"category":["Epigenetics Kits and Reagents","Epigenetics Kits and Reagents/CUTANA™ ChIC / CUT&RUN Assays"],"summary":"\n \n \n Type: Nuclease\n \n \n Mol Wgt: 43.7 kDa\n \n \n \n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/694/689/Screen_Shot_2020-02-12_at_11.01.55_AM__17144.1581530752.png?c=2","alt":"CUTANA™ pAG-MNase for ChIC/CUT&RUN Workflows"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/694/689/Screen_Shot_2020-02-12_at_11.01.55_AM__17144.1581530752.png?c=2","alt":"CUTANA™ pAG-MNase for ChIC/CUT&RUN Workflows"}],"date_added":"12th Aug 2019","pre_order":false,"show_cart_action":true,"has_options":true,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":1174,"name":"Internal Comment","value":"Excess in bottom of Venom"},{"id":1175,"name":"Internal Comment","value":"Bulk in Psylocke"}],"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"price":{"without_tax":{"currency":"USD","formatted":"$335.00","value":335},"price_range":{"min":{"without_tax":{"currency":"USD","formatted":"$335.00","value":335},"tax_label":"Sales Tax"},"max":{"without_tax":{"currency":"USD","formatted":"$1,295.00","value":1295},"tax_label":"Sales Tax"}},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=694"},{"id":889,"sku":null,"name":"CUTANA™ CUT&RUN Library Prep Kit","url":"https://www.epicypher.com/products/epigenetics-reagents-and-assays/cutana-cut-and-run-library-prep-kit","availability":"","rating":null,"brand":{"name":null},"category":["Epigenetics Kits and Reagents","Epigenetics Kits and Reagents/CUTANA™ ChIC / CUT&RUN Assays"],"summary":"\n \n \n \n \n \n \n \n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/889/997/Epicypher_CUTANA_CUTRUN_LibraryPrep_Box__64431.1646406521.png?c=2","alt":"CUTANA™ CUT&RUN Library Prep Kit"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/889/997/Epicypher_CUTANA_CUTRUN_LibraryPrep_Box__64431.1646406521.png?c=2","alt":"CUTANA™ CUT&RUN Library Prep Kit"}],"date_added":"31st Jan 2022","pre_order":false,"show_cart_action":true,"has_options":true,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":983,"name":"Pack Size","value":"48 Reactions"}],"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"price":{"without_tax":{"currency":"USD","formatted":"$1,525.00","value":1525},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=889"},{"id":906,"sku":"13-2021","name":"Menin CUTANA™ CUT&RUN Antibody","url":"https://www.epicypher.com/products/antibodies/cutana-cut-run-antibodies/cut-run-antibodies-chromatin-associated-proteins/menin-cutana-cut-run-antibody","availability":"","rating":null,"brand":{"name":null},"category":["Antibodies/CUTANA™ CUT&RUN Antibodies","Antibodies/CUTANA™ CUT&RUN Antibodies/CUT&RUN Antibodies - Chromatin-Associated Proteins","Epigenetics Kits and Reagents/CUTANA™ ChIC / CUT&RUN Assays"],"summary":"\n \n \n Type: Polyclonal\n \n \n Host: Rabbit\n \n \n Applications: CUT&R","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/906/1010/chd3-cutana-cutandrun-antibody__66975.1645734525__91145.1648666822__40012.1648670522.jpg?c=2","alt":"Menin CUTANA™ CUT&RUN Antibody"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/906/1010/chd3-cutana-cutandrun-antibody__66975.1645734525__91145.1648666822__40012.1648670522.jpg?c=2","alt":"Menin CUTANA™ CUT&RUN Antibody"}],"date_added":"30th Mar 2022","pre_order":false,"show_cart_action":true,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":1030,"name":"Pack Size","value":"100 µL"}],"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"price":{"without_tax":{"currency":"USD","formatted":"$495.00","value":495},"tax_label":"Sales Tax"},"out_of_stock_message":false,"add_to_wishlist_url":"/wishlist.php?action=add&product_id=906"},{"id":804,"sku":"13-2008","name":"CHD1 CUTANA™ CUT&RUN Antibody","url":"https://www.epicypher.com/products/antibodies/chd1-cutana-cut-run-antibody","availability":"","rating":null,"brand":{"name":null},"category":["Antibodies","Antibodies/CUTANA™ CUT&RUN Antibodies","Antibodies/CUTANA™ CUT&RUN Antibodies/CUT&RUN Antibodies - Chromatin-Associated Proteins","Epigenetics Kits and Reagents/CUTANA™ ChIC / CUT&RUN Assays"],"summary":"\n \n \n Type: Polyclonal\n \n \n Host: Rabbit\n \n \n Applications: CUT&RUN","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/804/793/2020.12.08_CUTANA_ChAPs_Antibody_thumbnail_RGB_LS__39482.1614276466.png?c=2","alt":"CHD1 CUTANA™ CUT&RUN Antibody"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/804/793/2020.12.08_CUTANA_ChAPs_Antibody_thumbnail_RGB_LS__39482.1614276466.png?c=2","alt":"CHD1 CUTANA™ CUT&RUN Antibody"}],"date_added":"28th Jan 2021","pre_order":false,"show_cart_action":true,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":666,"name":"Pack Size","value":"100 µL"}],"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"add_to_cart_url":"https://www.epicypher.com/cart.php?action=add&product_id=804","price":{"without_tax":{"currency":"USD","formatted":"$495.00","value":495},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=804"},{"id":909,"sku":"13-2026","name":"EZH2 CUTANA™ CUT&RUN Antibody","url":"https://www.epicypher.com/products/antibodies/cutana-cut-run-antibodies/cut-run-antibodies-chromatin-associated-proteins/ezh2-cutana-cut-run-antibody","availability":"","rating":null,"brand":{"name":null},"category":["Antibodies/CUTANA™ CUT&RUN Antibodies","Antibodies/CUTANA™ CUT&RUN Antibodies/CUT&RUN Antibodies - Chromatin-Associated Proteins","Epigenetics Kits and Reagents/CUTANA™ ChIC / CUT&RUN Assays"],"summary":"\n \n \n Type: Polyclonal\n \n \n Host: Rabbit\n \n \n Applications: CUT&R","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/909/1007/chd3-cutana-cutandrun-antibody__66975.1645734525__91145.1648666822__81008.1648670468.jpg?c=2","alt":"EZH2 CUTANA™ CUT&RUN Antibody"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/909/1007/chd3-cutana-cutandrun-antibody__66975.1645734525__91145.1648666822__81008.1648670468.jpg?c=2","alt":"EZH2 CUTANA™ CUT&RUN Antibody"}],"date_added":"30th Mar 2022","pre_order":false,"show_cart_action":true,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":1032,"name":"Pack Size","value":"100 µL"}],"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"add_to_cart_url":"https://www.epicypher.com/cart.php?action=add&product_id=909","price":{"without_tax":{"currency":"USD","formatted":"$495.00","value":495},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=909"}],"shipping_messages":[],"rating":0,"meta_keywords":"CUTANA, CUT&RUN, CUT&RUN kit, CUT and RUN, CUT and RUN kit, CUTANA CUT&RUN Kit, pAG-MNase kit, genomic mapping, ChIP-Seq","show_quantity_input":1,"title":"CUTANA™ ChIC/CUT&RUN Kit","gift_wrapping_available":false,"min_purchase_quantity":0,"customizations":[],"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/772/998/Epicypher_CUTANA_CUTRUN_Box__31908.1646406725.png?c=2","alt":"CUTANA™ ChIC/CUT&RUN Kit"}]} Pack size: 48 Reactions
Description
The CUTANA™ ChIC/CUT&RUN Kit enables streamlined chromatin profiling of histone post-translational modifications (PTMs) and chromatin associated proteins.
Save 10% when you buy with our Library Prep Kit for a streamlined CUT&RUN workflow! Learn more >>
The CUT&RUN Kit Version 4 (v4) now uses SPRI magnetic beads for DNA purification instead of DNA spin columns, enabling multi-channel pipetting throughout for increased throughput and reproducibility. Positive (H3K4me3) and negative (IgG) control antibodies are included to pair with SNAP-CUTANA™ spike-in controls for assay optimization and continuous assay monitoring (Figure 2). E. coli DNA is included for data normalization. The kit is compatible with a variety of inputs including cells or nuclei derived from native, cryopreserved, or cross-linked samples. While it is recommended to start with 500,000 cells, comparable data can be generated using as few as 5,000 cells. The inclusion of controls, as well as compatibility with diverse target types, sample inputs, and low cell numbers, make this kit the go-to solution for chromatin mapping experiments. To place bulk orders, contact info@epicypher.com.
Keep learning by checking out our Tech Support Center.
Validation Data
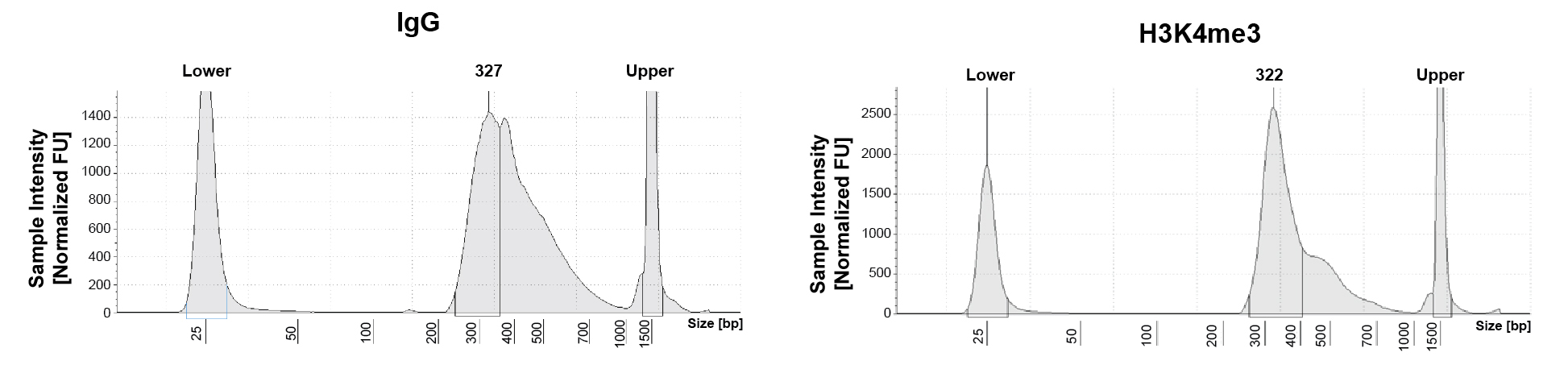
Figure 1: CUT&RUN DNA fragment size distribution
analysis
CUT&RUN was performed as described in
Figure 5. Library DNA was analyzed by
Agilent Tapestation®. This analysis
confirmed that mononucleosomes were predominantly
enriched in CUT&RUN (~300 bp peaks represent 150 bp
nucleosomes + sequencing adapters).
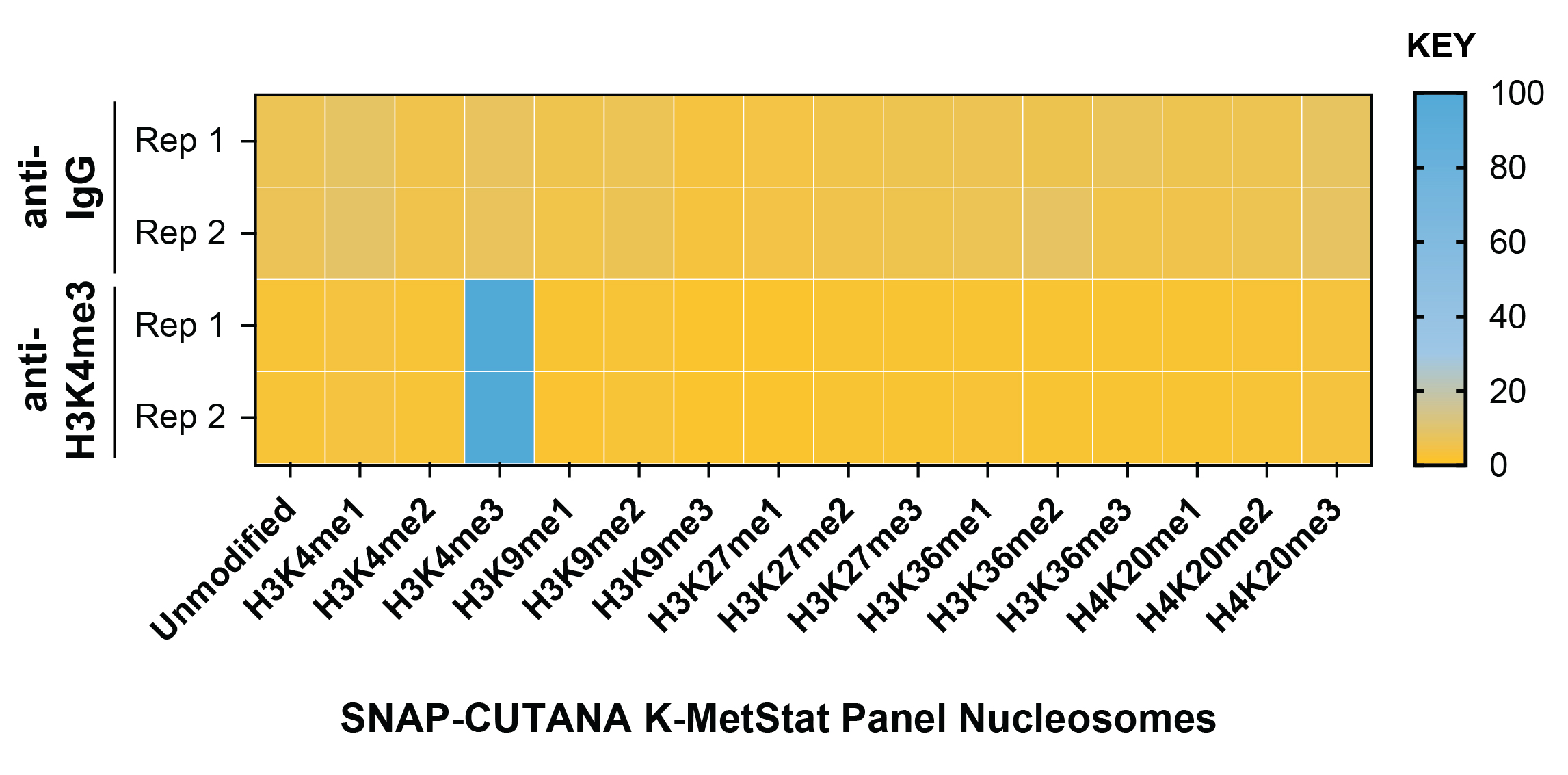
Figure 2: SNAP-CUTANA™ K-MetStat Spike-in
controls
DNA-barcoded designer nucleosomes (dNucs) representing
16 K-methyl PTMs: mono-, di-, and tri-methylation at
H3K4, H3K9, H3K27, H3K36, and H4K20, as well as
unmodified control, were spiked into CUT&RUN reactions
prior to the addition of antibodies (IgG, H3K4me3).
Instances of each spike-in barcode were counted and
normalized from raw fastq files using the shell script
and analysis excel sheet available on the spike-in
product page (EpiCypher
19-1002). Barcodes for IgG (top; normalized to total reads)
and H3K4me3 (bottom; normalized to on-target)
antibodies are shown. The spike-ins confirmed optimal
experimental conditions (H3K4me3 antibody specifically
recovered the target dNuc, while IgG showed no
preferential enrichment).
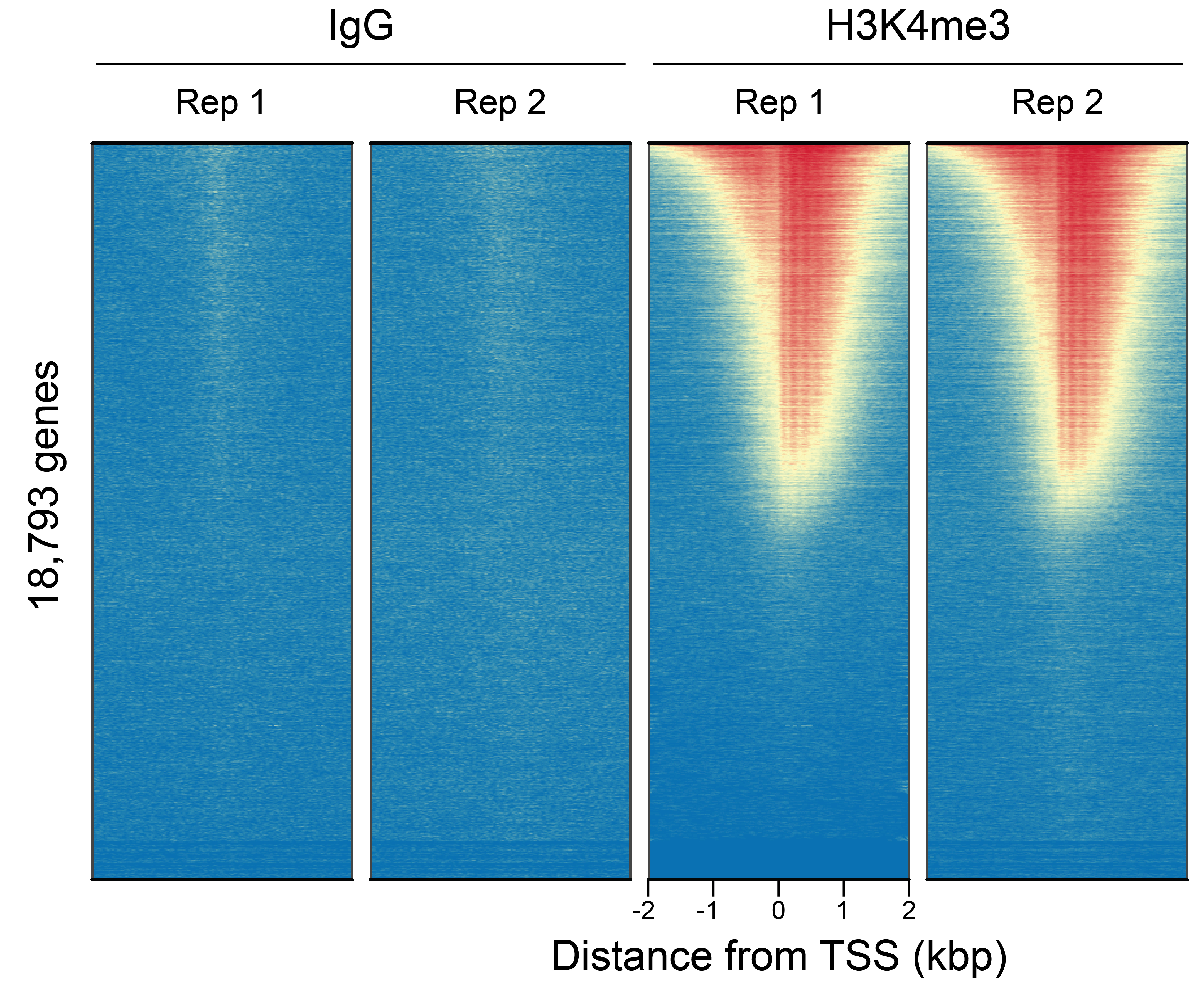
Figure 3: CUT&RUN genome-wide heatmaps
CUT&RUN was performed as described in
Figure 5. Heatmaps show two
replicates (“Rep”) of IgG (left) and H3K4me3 (right)
kit control antibodies in aligned rows ranked by
intensity (top to bottom) and colored such that red
indicates high localized enrichment and blue denotes
background signal.
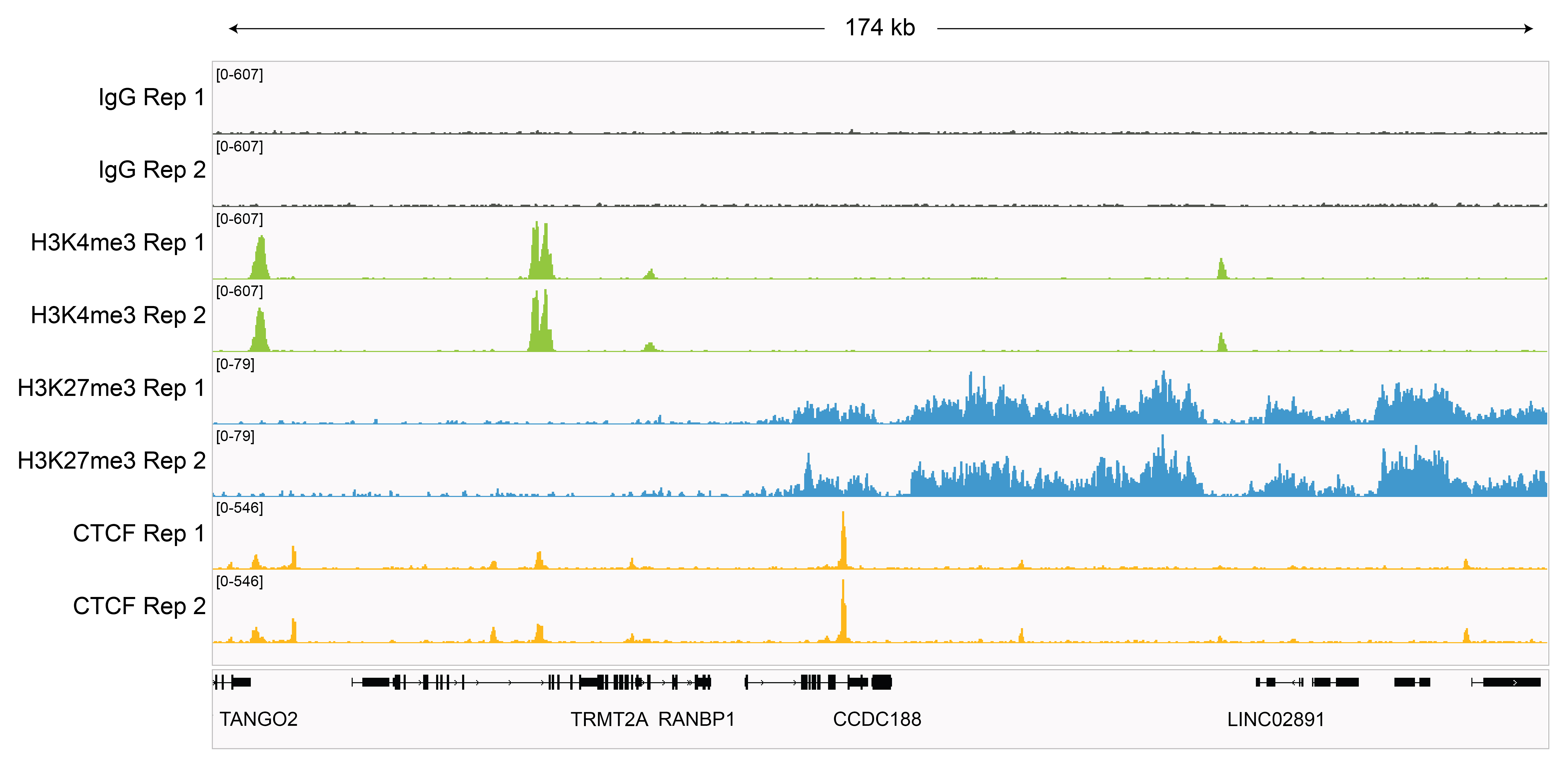
Figure 4: Representative gene browser
tracks
CUT&RUN was performed as described in
Figure 5. A representative 174 kb
window at the TRMT2A gene is shown for two replicates
(“Rep”) of IgG and H3K4me3 kit control antibodies.
Representative tracks are also shown for antibodies to
H3K27me3 and the transcription factor CTCF. The
CUT&RUN kit produced the expected genomic distribution
for each target. Images were generated using the
Integrative Genomics Viewer (IGV, Broad Institute).
Figure 5: CUT&RUN methods
CUT&RUN was performed using the CUTANA™ ChIC/CUT&RUN
Kit starting with 500k K562 cells with 0.5 µg of
either IgG (EpiCypher
13-0042), H3K4me3 (EpiCypher
13-0041), H3K27me3 (EpiCypher
13-0055), or 0.125 µg of CTCF (EpiCypher
13-2014) antibodies in duplicate. Library preparation was
performed with 5 ng of DNA (or the total amount
recovered if less than 5 ng) using the CUTANA™ Library
Prep Kit (EpiCypher
14-1001/14-1002). Libraries were run on an Illumina NextSeq2000 with
paired-end sequencing (2x50 bp). Sample sequencing
depth was 3.5 million reads (IgG Rep 1), 3.8 million
reads (IgG Rep 2), 4.7 million reads (H3K4me3 Rep 1),
6.9 million reads (H3K4me3 Rep 2), 6.6 million reads
(H3K27me3 Rep 1), 4.7 million reads (H3K27me3 Rep 2),
3.9 million reads (CTCF Rep 1) and 4.6 million reads
(CTCF Rep 2). Data were aligned to the hg19 genome
using Bowtie2. Data were filtered to remove
duplicates, multi-aligned reads, and ENCODE DAC Exclusion List
regions.
Kit Contents
| Item | Cat. No. |
|---|---|
| CUTANA™ pAG-MNase for ChIC/CUT&RUN Workflows | 15-1016 |
| H3K4me3 Antibody, SNAP-Certified™ for CUT&RUN | 13-0041k |
| CUTANA™ Rabbit IgG CUT&RUN Negative Control Antibody | 13-0042k |
| SNAP-CUTANA™ K-MetStat Panel | 19-1002k |
| CUTANA™ Concanavalin A Conjugated Paramagnetic Beads | 21-1401 |
| CUTANA™ E. coli Spike-in DNA | 18-1401 |
| CUTANA™ CUT&RUN 8-Strip 0.2 mL Tubes | 10-0009a |
| Bead Activation Buffer | 21-1001 |
| Pre-Wash Buffer | 21-1002 |
| Stop Buffer | 21-1003 |
| 5% Digitonin | 21-1004k |
| 1 M Spermidine | 21-1005 |
| 0.5 M EDTA | 21-1006 |
| 100 mM Calcium Chloride | 21-1007 |
| 0.1X TE Buffer | 21-1025 |
| SPRIselect reagent manufactured by Beckman Coulter, Inc. | 21-1405 |
Recommended Accessory Products
| Item | Cat. No. |
|---|---|
| CUTANA™ CUT&RUN Library Prep Kit | 14-1001/14-1002 |
| SNAP-CUTANA™ K-MetStat Panel | 19-1002 |
| CUT&RUN Antibodies | See the list |
| Magnetic Separation Rack, 0.2 mL Tubes | 10-0008 |
| Magnetic Separation Rack, 1.5 mL Tubes | 10-0012 |
| CUTANA™ Nuclei Extraction Buffer | 21-1026 |
Technical Information
Additional Information
Beckman Coulter, the stylized logo, and SPRIselect are trademarks or registered trademarks of Beckman Coulter, Inc. in the United States and other countries.