Histone H3K27me3 Antibody, SNAP-ChIP® Certified, CUTANA™ CUT&RUN Compatible *DISCONTINUED*
SKU: 13-0030
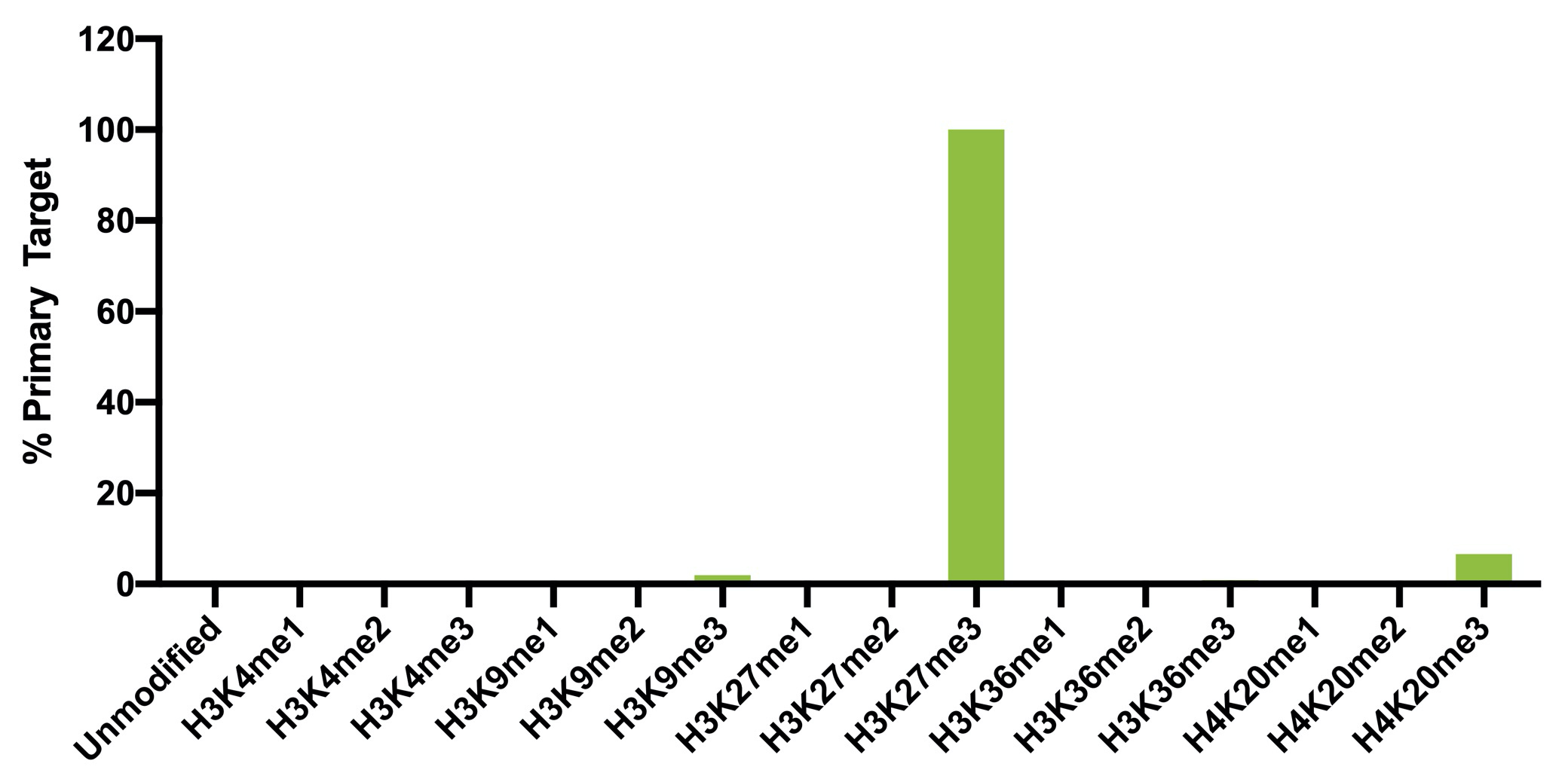
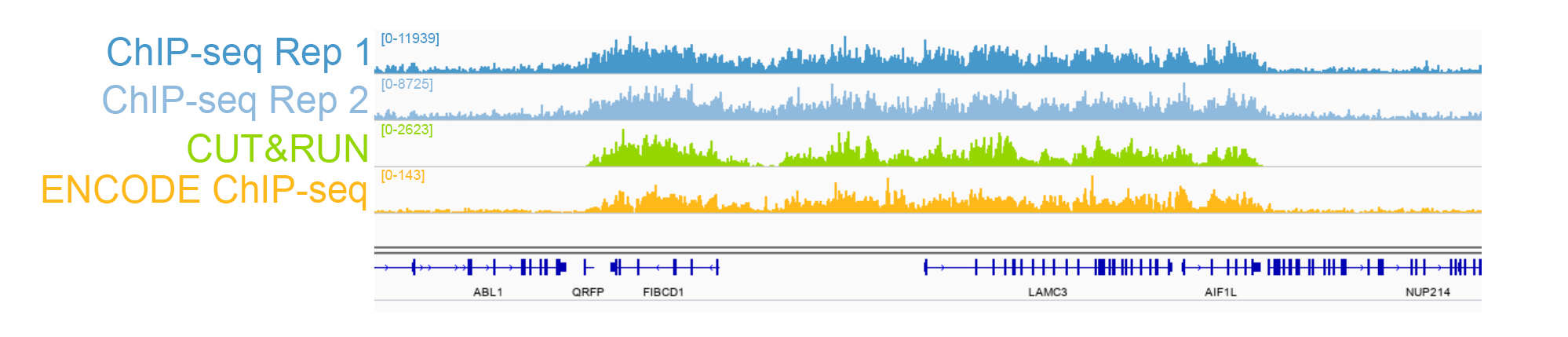
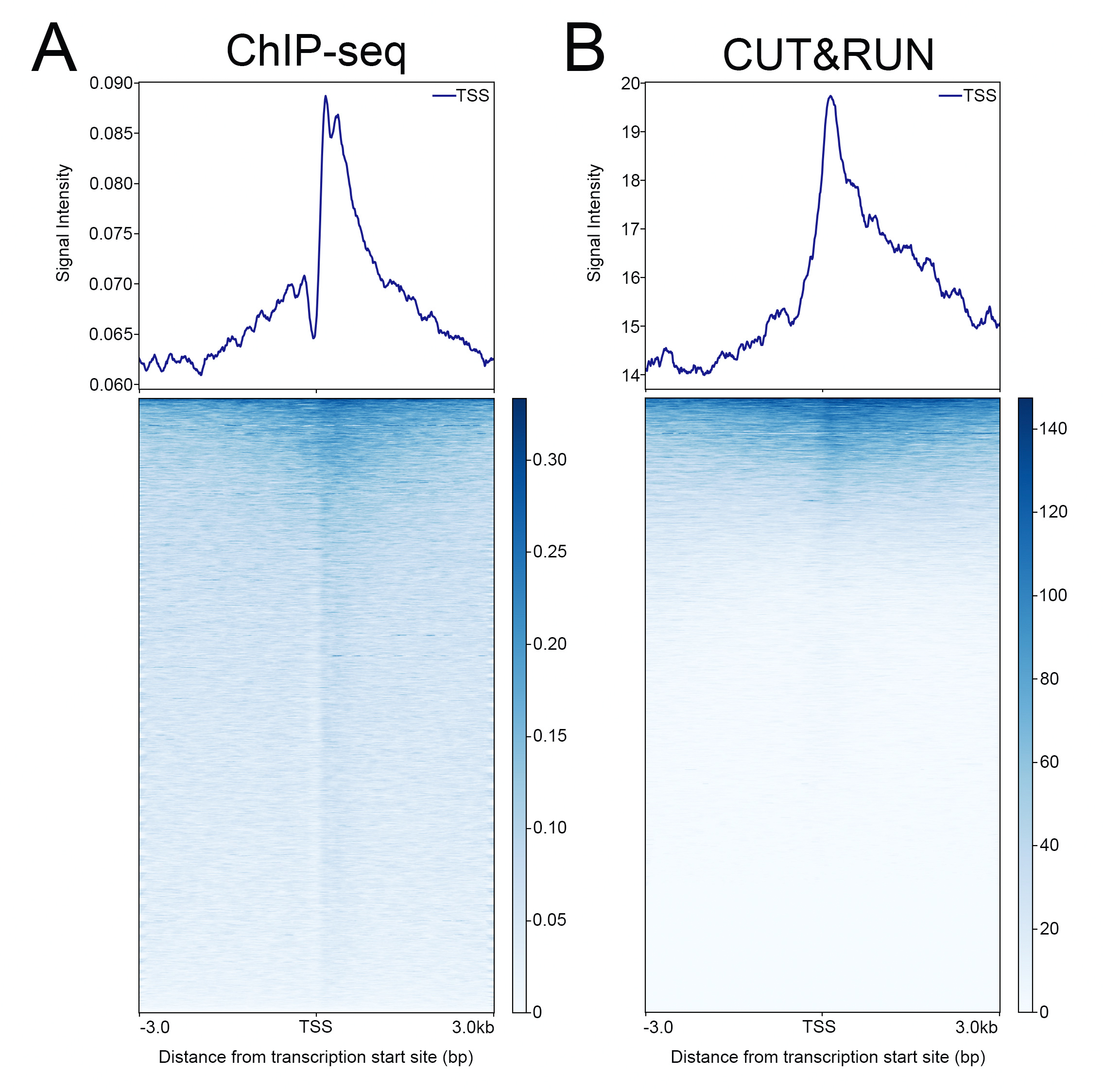
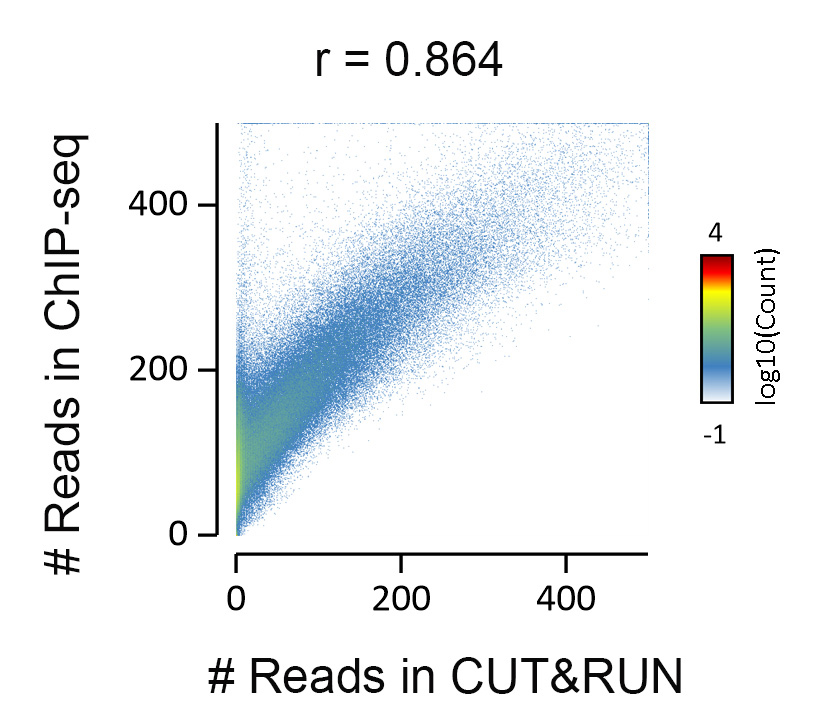
{"url":"https://www.epicypher.com/products/antibodies/snap-chip-certified-antibodies/histone-h3k27me3-antibody-snap-chip-certified-cutana-cut-run-compatible-discontinued","add_this":[{"service":"facebook","annotation":""},{"service":"email","annotation":""},{"service":"print","annotation":""},{"service":"twitter","annotation":""},{"service":"linkedin","annotation":""}],"warranty":"","gtin":"","max_purchase_quantity":0,"id":"628","can_purchase":false,"meta_description":"Histone H3K27me3 Antibody, SNAP-ChIP® Certified, CUTANA™ CUT&RUN Compatible (13-0030)","category":[],"meta_keywords":"CUTANA™ Compatible, Histone H3K27me3 Antibody - SNAP ChIP® Certified, 13-0030, H3K27me3, SNAP, H3K27me3 SNAP ChIP, SNAP ChIP, H3K27me3 SNAP-ChIP, SNAP-ChIP, H3K27me3 SNAP, H3K27me3 Antibody, CUT&RUN, cut and run, cleavage under targets and release using nuclease, ChIC, 13-0030 ","AddThisServiceButtonMeta":"","images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/628/940/histone-h3k27me3-antibody-snap-chip-certified-cutana-cutandrun-compatible-discontinued__17318.1645734531.jpg?c=2","alt":"Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUTandRUN Compatible - DISCONTINUED"}],"main_image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/628/940/histone-h3k27me3-antibody-snap-chip-certified-cutana-cutandrun-compatible-discontinued__17318.1645734531.jpg?c=2","alt":"Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUTandRUN Compatible - DISCONTINUED"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=628","shipping":[],"num_reviews":0,"weight":"0.00 LBS","custom_fields":[{"id":"470","name":"Pack Size","value":"100 μg"}],"sku":"13-0030","description":"<table border=\"0\" cellpadding=\"2\" cellspacing=\"2\" style=\"width: 100%;\"> <tbody> <tr valign=\"top\"> \n <a\n style=\"color: #fff\"\n href=\"/products/antibodies/h3k27me3-antibody-snap-certified-for-cut-run-and-cut-and-tag\">\n <p\n style=\"\n background-color: #4698cb;\n color: #fff;\n padding: 1.3rem;\n text-align: center;\n border-radius: 12px;\n margin-top: 2.5rem;\n \">\n Product Discontinued - see 13-0055 as a recommended alternative\n </p>\n </a>\n <td> <table border=\"0\" cellpadding=\"1\" cellspacing=\"1\" style=\"width: 95%;\"> <tbody> <tr> <td> <table border=\"0\" cellpadding=\"0\" cellspacing=\"0\" style=\"width: 100%;\"> <tbody> <tr> <td align=\"left\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Type: </strong>Monoclonal</span></td> \n <td align=\"left\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Host: </strong>Rabbit</span></td> </tr> <tr> <td align=\"left\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Mol Wgt.: </strong>15 kDa</span></td> <td align=\"left\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Appl.: </strong>ChIP, ChIP-Seq, CUT&RUN, Luminex, IF, IHC, WB</span></td> \n </tr> <tr> <td align=\"left\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Format: </strong>Aff. Pur. IgG</span></td> \n <td align=\"left\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Reactivity: </strong>H, M, WR</span></td> \n </tr> </tbody> </table> </td> </tr> \n \n <tr> <td valign=\"top\"> \n \n <span style=\"font-family:arial,helvetica,sans-serif;\"> <br /><p>This antibody has been discontinued. For alternative antibody options, please visit <a target=\"_blank\" rel=\"nofollow noreferrer noopener\" href=\"https://www.epicypher.com/snap-chip-certified-antibodies/\">https://www.epicypher.com/snap-chip-certified-antibodies/</a>.</p></span><br />\n \n <span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUT&RUN Compatible Description:</strong> This antibody meets EpiCypher’s “SNAP-ChIP® Certified” criteria for specificity and target enrichment in ChIP (5% recovery of target input determined using SNAP-ChIP K-MetStat Panel spike-in controls; EpiCypher Catalog No. 19-1001). Although its specificity in CUT&RUN has yet to be empirically determined <i>in situ</i> using spike-in controls, CUT&RUN data produced by this antibody shows a genome wide enrichment pattern characteristic of H3K27me3 and is highly correlated with ChIP-seq (Figures 3-5).</span><br /> \n </td> </tr> \n \n \n \n <tr> <td valign=\"top\"> <span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUT&RUN Compatible Immunogen:</strong></span> <span style=\"font-family:arial,helvetica,sans-serif;\"></span>A synthetic peptide corresponding to histone H3 trimethylated at lysine 27.<br />\n </p></td></tr> \n \n \n \n <tr> <td valign=\"top\"> <span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUT&RUN Compatible Formulation:</strong></span> <span style=\"font-family:arial,helvetica,sans-serif;\"></span>Protein A affinity-purified antibody (1 mg/mL) in PBS, with 0.09% sodium azide, 1% BSA, and 50% glycerol.<br />\n </p></td></tr>\n \n <tr>\n <td valign=\"top\"> <span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUT&RUN Compatible Storage and Stability:</strong></span> <span style=\"font-family:arial,helvetica,sans-serif;\">Stable for 1 years at -20°C from date of receipt.</span><br />\n </p></td></tr>\n \n <tr>\n <td valign=\"top\"> <span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUT&RUN Compatible Application Notes</strong></span> <span style=\"font-family:arial,helvetica,sans-serif;\"><span style=\"font-weight: normal;\"><strong>Recommended dilutions:</strong><br /> \n </span></span>ChIP/ChIP-seq: 2 - 5 μg per 1x10<sup>6</sup> cells CUT&RUN: 1:100 Luminex: 1:4000 IF/IHC: 0.5 - 2 μg/mL WB: 1 - 2 μg/mL</td></tr>\n \n <tr>\n <td valign=\"top\"> <span style=\"font-family:arial,helvetica,sans-serif;\"><span style=\"font-size: 12pt;\"><strong>Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUT&RUN Compatible References:</strong> <br/>Grzybowski\tet\tal\t(2015)\tMol\tCell\t58:886<br/>\n Shah\tet\tal\t(2018)\tMol\tCell\t72:162</span></span></p></td></tr>\n \n <tr>\n <td valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\">View technical datasheet for this product. <a href=\"https://www.epicypher.com/content/documents/tds/13-0030.pdf\" target=\"_new\"> <img alt=\"13-0030 Datasheet\" height=\"40\" src=\"https://cdn7.bigcommerce.com/s-y9o92/content/documents/tds/icon.png\" width=\"30\" /></a></span></td> \n </tr> <tr> <td> <p></p> <p><span style=\"font-family:arial,helvetica,sans-serif;\"><span style=\"font-size: 10pt;\"><strong>Applications Key: ChIP</strong>-Chromatin IP; <strong>E</strong>-ELISA; <strong>FACS</strong>-Flow cytometry; <strong>IF</strong>-Immunofluorescence; <strong>IHC</strong>-Immunohistochemistry; <strong>IP</strong>-Immunoprecipitation; <strong>WB</strong>-Western Blotting</span></span></p> <p><span style=\"font-family:arial,helvetica,sans-serif;\"><span style=\"font-size: 10pt;\"><strong>Reactivity Key: </strong><strong>B</strong>-Bovine; <strong>Ce</strong>-C. elegans; <strong>Ch</strong>-Chicken; <strong>Dm</strong>- Drosophila; <strong>Eu</strong>-Eukaryote; <strong>H</strong>-Human; <strong>M</strong>-Mouse; <strong>Ma</strong>-Mammal; <strong>R</strong>-Rat; <strong>Sc</strong>-S.cerevesiae; <strong>Sp</strong>-S. pombe; <strong>WR</strong>-Wide Range (predicted); <strong>X</strong>-Xenopus; <strong>Z</strong>-Zebrafish</span></span></p> </td> </tr> </tbody> </table> </td>\n \n <td><table border=\"0\" cellpadding=\"3\" cellspacing=\"3\" style=\"width: 300px;\"> <tbody> <tr> <td align=\"center\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"https://www.epicypher.com/content/images/products/antibodies/13_0030_luminex_tds.jpg\" target=\"_new\"><img alt=\"13-0030 Luminex Data\" height=\"100\" src=\"https://cdn7.bigcommerce.com/s-y9o92/content/images/products/antibodies/13_0030_luminex_tds.jpg\" width=\"150\" /></a></span></td> </tr> <tr> <td align=\"left\" valign=\"top\"> <p><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Luminex multiplexed specificity profiling:</strong> H3K27me3 antibody was assessed using a Luminex® based approach employing dCypher™ Nucleosome K-MetStat Panel (EpiCypher Catalog No. 16-9002). The panel comprises biotinylated designer nucleosomes (x-axis) individually coupled to color coded Luminex Magplex® beads. Antibody binding to the panel of 16 nucleosomes was tested in multiplex at a 1:4000 dilution, and detected with second layer anti-IgG*PE. Data was generated using a Luminex FlexMAP3D®. Data is normalized to target (H3K27me3; set to 100).<br /> <strong>(Click image to enlarge) </strong></span></p> <p></p> </td> </tr> \n \n <tr valign=\"top\"> <td align=\"center\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"https://www.epicypher.com/content/images/products/antibodies/13_0030_snap_tds.jpg\" target=\"_new\"><img alt=\"13-0030 SNAP-ChIP data\" height=\"30\" src=\"https://cdn11.bigcommerce.com/s-y9o92/content/images/products/antibodies/13_0030_snap_tds.jpg\" width=\"50\" /></a></span></td> </tr> <tr> <td align=\"left\" valign=\"top\"> <p><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>SNAP-ChIP-qPCR specificity and enrichment analysis:</strong> H3K27me3 antibody (5 μg) was tested in a native ChIP experiment using chromatin from K-562 cells (5 μg) with the SNAP-ChIP K-MetStat Panel (EpiCypher Catalog No. 19-1001) spiked-in prior to micrococcal nuclease digestion. Specificity (left y-axis) was determined by qPCR for the DNA barcodes corresponding to modified nucleosomes in the SNAP-ChIP panel (x-axis). Black bar represents antibody efficiency (right y-axis; log scale) and indicates percentage of the target immunoprecipitated relative to input. Error bars represent mean ± SEM in replicate ChIP experiments.<br /> <strong>(Click image to enlarge) </strong></span></p> <p></p> </td> </tr> \n \n <tr valign=\"top\"> <td align=\"center\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"https://www.epicypher.com/content/images/products/antibodies/13_0030_seq_tracks.jpg\" target=\"_new\"><img alt=\"13-0030 Sequencing Tracks\" height=\"100\" src=\"https://cdn11.bigcommerce.com/s-y9o92/content/images/products/antibodies/13_0030_seq_tracks.jpg\" width=\"127\" /></a></span></td> </tr> <tr> <td align=\"left\" valign=\"top\"> <p><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>H3K27me3 SNAP-ChIP-seq and CUT&RUN representative tracks: </strong>A gene browser shot generated using the Integrative Genomics Viewer (IGV, Broad Institute) shows a representative locus for EpiCypher H3K27me3 ChIP-seq replicate experiments (blue tracks, 5 μg antibody) and CUT&RUN (green track, 1:100 antibody dilution). For comparison ENCODE H3K27me3 ChIP-seq using a different antibody is shown (bottom orange track, GEO accession number GSM733658). Similar results in peak structure and location were observed throughout the genome for EpiCypher H3K27me3 antibody in ChIP-seq and CUT&RUN. <i>Methods: Native ChIP-seq was performed as described (Shah et al., Mol Cell 2018). CUT&RUN was performed using EpiCypher CUTANA pAG-MNase for ChIC/CUT&RUN (EpiCypher Catalog No. 15-1016) as described (EpiCypher.com/cutana-protocol). Library preparation was performed with 10 ng DNA using the NEBNext® UltraTM II DNA Library Prep Kit for Illumina®. ChIP libraries were sequenced on an Illumina NextSeq 550 (2x150bp paired end). The total number of reads was 37.8 and 34.9 million for the ChIP-seq replicates and 8.0 million for CUT&RUN.</i><br /> <strong>(Click image to enlarge) </strong></span></p> <p></p> </td> </tr>\n \n <tr valign=\"top\"> <td align=\"center\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"https://www.epicypher.com/content/images/products/antibodies/13_0030_tss.jpg\" target=\"_new\"><img alt=\"13-0030 Genome Wide Analysis\" height=\"100\" src=\"https://cdn11.bigcommerce.com/s-y9o92/content/images/products/antibodies/13_0030_tss.jpg\" width=\"127\" /></a></span></td> </tr> <tr> <td align=\"left\" valign=\"top\"> <p><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>ChIP-seq and CUT&RUN genome wide analysis: </strong>EpiCypher H3K27me3 antibody was tested in native ChIP-seq <strong> A </strong> and CUT&RUN <strong> B </strong> using the methods described above. Genome-wide analysis of H3K27me3 enrichment (signal intensity) flanking annotated transcription start sits (TSSs; +/- 3kb) is graphed as a cumulative histogram plot (top) and shown in a heatmap (bottom). Individual gene loci in each row of the heatmap are colored by signal intensity and sorted by strongest to lowest enrichment (top to bottom). EpiCypher H3K27me3 an2body displays a characteristic enrichment pattern downstream of the TSS, remaining elevated throughout the body of the gene.<br /> <strong>(Click image to enlarge) </strong></span></p> <p></p> </td> </tr> \n \n <tr valign=\"top\"> <td align=\"center\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"https://www.epicypher.com/content/images/products/antibodies/13_0030_ca_tracks.jpg\" target=\"_new\"><img alt=\"13-0030 Correlation Analysis\" height=\"100\" src=\"https://cdn11.bigcommerce.com/s-y9o92/content/images/products/antibodies/13_0030_ca_tracks.jpg\" width=\"127\" /></a></span></td> </tr> <tr> <td align=\"left\" valign=\"top\"> <p><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>ChIP-seq vs. CUT&RUN correlation analysis: </strong>Genome-wide correlation analysis was performed to compare EpiCypher H3K27me3 antibody enrichment in ChIP-seq and CUT&RUN. The number of reads per 10 kb binned region across the genome is plotted for CUT&RUN (x-axis) vs. ChIP-seq (y-axis) (EaSeq). ChIP-seq and CUT&RUN data generated using this antibody are highly correlated (Pearson correlation r = 0.864).<br /> <strong>(Click image to enlarge) </strong></span></p> <p></p> </td> </tr> \n \n <tr valign=\"top\"> <td align=\"center\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"https://www.epicypher.com/content/images/products/antibodies/13_0030_wb_tds.jpg\" target=\"_new\"><img alt=\"13-0030 Western Blot\" height=\"100\" src=\"https://cdn11.bigcommerce.com/s-y9o92/content/images/products/antibodies/13_0030_wb_tds.jpg\" width=\"127\" /></a></span></td> </tr> <tr> <td align=\"left\" valign=\"top\"> <p><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Western Blot Analysis: </strong>Recombinant histone H3.3 (Lane 1) and acid extracts of HeLa cells (Lane 2) were blotted onto PVDF and probed with 1 µg/mL EpiCypher H3K27me3 Antibody.<br /> <strong>(Click image to enlarge) </strong></span></p> <p></p> </td> </tr>\n \n <tr valign=\"top\"> <td align=\"center\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"https://www.epicypher.com/content/images/products/antibodies/13_0030_IHC_tds.jpg\" target=\"_new\"><img alt=\"13-0030 Immunohistochemistry\" height=\"100\" src=\"https://cdn11.bigcommerce.com/s-y9o92/content/images/products/antibodies/13_0030_IHC_tds.jpg\" width=\"127\" /></a></span></td> </tr> <tr> <td align=\"left\" valign=\"top\"> <p><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Immunohistochemistry (IHC): </strong>IHC staining of HepG2 cells using EpiCypher\n H3K27me3\tAntibody.<br /> <strong>(Click image to enlarge) </strong></span></p> <p></p> </td> </tr></tbody> </table> </td> \n </tr> </tbody> </table> \n \n <style>\n .button-group {\n display: none;\n }\n </style>","tags":[],"detail_messages":"","availability":"","page_title":"Histone H3K27me3 Antibody, SNAP-ChIP® Certified, CUTANA™ CUT&RUN Compatible (13-0030)","mpn":"","upc":null,"options":[],"related_products":[{"id":411,"sku":null,"name":"SNAP-ChIP® K-MetStat Panel","url":"https://www.epicypher.com/products/nucleosomes/snap-chip-k-metstat-panel","availability":"","rating":null,"brand":{"name":null},"category":["Nucleosomes","Nucleosomes/SNAP-ChIP<sup>®</sup> Spike-ins"],"summary":"\n\t\n\t\t\n\t\t\t\n\t\t\t\n\t\t\t\t\n\t\t\t\t\t\n\t\t\t\t\t\t\n\t\t\t\t\t\t\n\n\t\t\t\t\t\t\n\t\t\t\t\t\tSNAP-ChIP K-MetStat Panel Description: A panel ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/411/282/SNAP_Chip__53961.1569012494.png?c=2","alt":"SNAP-ChIP® K-MetStat Panel"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/411/282/SNAP_Chip__53961.1569012494.png?c=2","alt":"SNAP-ChIP® K-MetStat Panel"}],"date_added":"18th Sep 2017","pre_order":false,"show_cart_action":true,"has_options":true,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":null,"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"price":{"without_tax":{"currency":"USD","formatted":"$365.00","value":365},"price_range":{"min":{"without_tax":{"currency":"USD","formatted":"$365.00","value":365},"tax_label":"Sales Tax"},"max":{"without_tax":{"currency":"USD","formatted":"$2,795.00","value":2795},"tax_label":"Sales Tax"}},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=411"}],"shipping_messages":[],"rating":0,"title":"Histone H3K27me3 Antibody, SNAP-ChIP® Certified, CUTANA™ CUT&RUN Compatible *DISCONTINUED*","gift_wrapping_available":false,"min_purchase_quantity":0} Pack Size: 100 μg
{"url":"https://www.epicypher.com/products/antibodies/snap-chip-certified-antibodies/histone-h3k27me3-antibody-snap-chip-certified-cutana-cut-run-compatible-discontinued","add_this":[{"service":"facebook","annotation":""},{"service":"email","annotation":""},{"service":"print","annotation":""},{"service":"twitter","annotation":""},{"service":"linkedin","annotation":""}],"warranty":"","gtin":"","max_purchase_quantity":0,"id":"628","can_purchase":false,"meta_description":"Histone H3K27me3 Antibody, SNAP-ChIP® Certified, CUTANA™ CUT&RUN Compatible (13-0030)","category":[],"meta_keywords":"CUTANA™ Compatible, Histone H3K27me3 Antibody - SNAP ChIP® Certified, 13-0030, H3K27me3, SNAP, H3K27me3 SNAP ChIP, SNAP ChIP, H3K27me3 SNAP-ChIP, SNAP-ChIP, H3K27me3 SNAP, H3K27me3 Antibody, CUT&RUN, cut and run, cleavage under targets and release using nuclease, ChIC, 13-0030 ","AddThisServiceButtonMeta":"","images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/628/940/histone-h3k27me3-antibody-snap-chip-certified-cutana-cutandrun-compatible-discontinued__17318.1645734531.jpg?c=2","alt":"Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUTandRUN Compatible - DISCONTINUED"}],"main_image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/628/940/histone-h3k27me3-antibody-snap-chip-certified-cutana-cutandrun-compatible-discontinued__17318.1645734531.jpg?c=2","alt":"Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUTandRUN Compatible - DISCONTINUED"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=628","shipping":[],"num_reviews":0,"weight":"0.00 LBS","custom_fields":[{"id":"470","name":"Pack Size","value":"100 μg"}],"sku":"13-0030","description":"<table border=\"0\" cellpadding=\"2\" cellspacing=\"2\" style=\"width: 100%;\"> <tbody> <tr valign=\"top\"> \n <a\n style=\"color: #fff\"\n href=\"/products/antibodies/h3k27me3-antibody-snap-certified-for-cut-run-and-cut-and-tag\">\n <p\n style=\"\n background-color: #4698cb;\n color: #fff;\n padding: 1.3rem;\n text-align: center;\n border-radius: 12px;\n margin-top: 2.5rem;\n \">\n Product Discontinued - see 13-0055 as a recommended alternative\n </p>\n </a>\n <td> <table border=\"0\" cellpadding=\"1\" cellspacing=\"1\" style=\"width: 95%;\"> <tbody> <tr> <td> <table border=\"0\" cellpadding=\"0\" cellspacing=\"0\" style=\"width: 100%;\"> <tbody> <tr> <td align=\"left\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Type: </strong>Monoclonal</span></td> \n <td align=\"left\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Host: </strong>Rabbit</span></td> </tr> <tr> <td align=\"left\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Mol Wgt.: </strong>15 kDa</span></td> <td align=\"left\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Appl.: </strong>ChIP, ChIP-Seq, CUT&RUN, Luminex, IF, IHC, WB</span></td> \n </tr> <tr> <td align=\"left\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Format: </strong>Aff. Pur. IgG</span></td> \n <td align=\"left\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Reactivity: </strong>H, M, WR</span></td> \n </tr> </tbody> </table> </td> </tr> \n \n <tr> <td valign=\"top\"> \n \n <span style=\"font-family:arial,helvetica,sans-serif;\"> <br /><p>This antibody has been discontinued. For alternative antibody options, please visit <a target=\"_blank\" rel=\"nofollow noreferrer noopener\" href=\"https://www.epicypher.com/snap-chip-certified-antibodies/\">https://www.epicypher.com/snap-chip-certified-antibodies/</a>.</p></span><br />\n \n <span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUT&RUN Compatible Description:</strong> This antibody meets EpiCypher’s “SNAP-ChIP® Certified” criteria for specificity and target enrichment in ChIP (5% recovery of target input determined using SNAP-ChIP K-MetStat Panel spike-in controls; EpiCypher Catalog No. 19-1001). Although its specificity in CUT&RUN has yet to be empirically determined <i>in situ</i> using spike-in controls, CUT&RUN data produced by this antibody shows a genome wide enrichment pattern characteristic of H3K27me3 and is highly correlated with ChIP-seq (Figures 3-5).</span><br /> \n </td> </tr> \n \n \n \n <tr> <td valign=\"top\"> <span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUT&RUN Compatible Immunogen:</strong></span> <span style=\"font-family:arial,helvetica,sans-serif;\"></span>A synthetic peptide corresponding to histone H3 trimethylated at lysine 27.<br />\n </p></td></tr> \n \n \n \n <tr> <td valign=\"top\"> <span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUT&RUN Compatible Formulation:</strong></span> <span style=\"font-family:arial,helvetica,sans-serif;\"></span>Protein A affinity-purified antibody (1 mg/mL) in PBS, with 0.09% sodium azide, 1% BSA, and 50% glycerol.<br />\n </p></td></tr>\n \n <tr>\n <td valign=\"top\"> <span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUT&RUN Compatible Storage and Stability:</strong></span> <span style=\"font-family:arial,helvetica,sans-serif;\">Stable for 1 years at -20°C from date of receipt.</span><br />\n </p></td></tr>\n \n <tr>\n <td valign=\"top\"> <span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUT&RUN Compatible Application Notes</strong></span> <span style=\"font-family:arial,helvetica,sans-serif;\"><span style=\"font-weight: normal;\"><strong>Recommended dilutions:</strong><br /> \n </span></span>ChIP/ChIP-seq: 2 - 5 μg per 1x10<sup>6</sup> cells CUT&RUN: 1:100 Luminex: 1:4000 IF/IHC: 0.5 - 2 μg/mL WB: 1 - 2 μg/mL</td></tr>\n \n <tr>\n <td valign=\"top\"> <span style=\"font-family:arial,helvetica,sans-serif;\"><span style=\"font-size: 12pt;\"><strong>Histone H3K27me3 Antibody, SNAP-ChIP Certified, CUTANA CUT&RUN Compatible References:</strong> <br/>Grzybowski\tet\tal\t(2015)\tMol\tCell\t58:886<br/>\n Shah\tet\tal\t(2018)\tMol\tCell\t72:162</span></span></p></td></tr>\n \n <tr>\n <td valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\">View technical datasheet for this product. <a href=\"https://www.epicypher.com/content/documents/tds/13-0030.pdf\" target=\"_new\"> <img alt=\"13-0030 Datasheet\" height=\"40\" src=\"https://cdn7.bigcommerce.com/s-y9o92/content/documents/tds/icon.png\" width=\"30\" /></a></span></td> \n </tr> <tr> <td> <p></p> <p><span style=\"font-family:arial,helvetica,sans-serif;\"><span style=\"font-size: 10pt;\"><strong>Applications Key: ChIP</strong>-Chromatin IP; <strong>E</strong>-ELISA; <strong>FACS</strong>-Flow cytometry; <strong>IF</strong>-Immunofluorescence; <strong>IHC</strong>-Immunohistochemistry; <strong>IP</strong>-Immunoprecipitation; <strong>WB</strong>-Western Blotting</span></span></p> <p><span style=\"font-family:arial,helvetica,sans-serif;\"><span style=\"font-size: 10pt;\"><strong>Reactivity Key: </strong><strong>B</strong>-Bovine; <strong>Ce</strong>-C. elegans; <strong>Ch</strong>-Chicken; <strong>Dm</strong>- Drosophila; <strong>Eu</strong>-Eukaryote; <strong>H</strong>-Human; <strong>M</strong>-Mouse; <strong>Ma</strong>-Mammal; <strong>R</strong>-Rat; <strong>Sc</strong>-S.cerevesiae; <strong>Sp</strong>-S. pombe; <strong>WR</strong>-Wide Range (predicted); <strong>X</strong>-Xenopus; <strong>Z</strong>-Zebrafish</span></span></p> </td> </tr> </tbody> </table> </td>\n \n <td><table border=\"0\" cellpadding=\"3\" cellspacing=\"3\" style=\"width: 300px;\"> <tbody> <tr> <td align=\"center\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"https://www.epicypher.com/content/images/products/antibodies/13_0030_luminex_tds.jpg\" target=\"_new\"><img alt=\"13-0030 Luminex Data\" height=\"100\" src=\"https://cdn7.bigcommerce.com/s-y9o92/content/images/products/antibodies/13_0030_luminex_tds.jpg\" width=\"150\" /></a></span></td> </tr> <tr> <td align=\"left\" valign=\"top\"> <p><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Luminex multiplexed specificity profiling:</strong> H3K27me3 antibody was assessed using a Luminex® based approach employing dCypher™ Nucleosome K-MetStat Panel (EpiCypher Catalog No. 16-9002). The panel comprises biotinylated designer nucleosomes (x-axis) individually coupled to color coded Luminex Magplex® beads. Antibody binding to the panel of 16 nucleosomes was tested in multiplex at a 1:4000 dilution, and detected with second layer anti-IgG*PE. Data was generated using a Luminex FlexMAP3D®. Data is normalized to target (H3K27me3; set to 100).<br /> <strong>(Click image to enlarge) </strong></span></p> <p></p> </td> </tr> \n \n <tr valign=\"top\"> <td align=\"center\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"https://www.epicypher.com/content/images/products/antibodies/13_0030_snap_tds.jpg\" target=\"_new\"><img alt=\"13-0030 SNAP-ChIP data\" height=\"30\" src=\"https://cdn11.bigcommerce.com/s-y9o92/content/images/products/antibodies/13_0030_snap_tds.jpg\" width=\"50\" /></a></span></td> </tr> <tr> <td align=\"left\" valign=\"top\"> <p><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>SNAP-ChIP-qPCR specificity and enrichment analysis:</strong> H3K27me3 antibody (5 μg) was tested in a native ChIP experiment using chromatin from K-562 cells (5 μg) with the SNAP-ChIP K-MetStat Panel (EpiCypher Catalog No. 19-1001) spiked-in prior to micrococcal nuclease digestion. Specificity (left y-axis) was determined by qPCR for the DNA barcodes corresponding to modified nucleosomes in the SNAP-ChIP panel (x-axis). Black bar represents antibody efficiency (right y-axis; log scale) and indicates percentage of the target immunoprecipitated relative to input. Error bars represent mean ± SEM in replicate ChIP experiments.<br /> <strong>(Click image to enlarge) </strong></span></p> <p></p> </td> </tr> \n \n <tr valign=\"top\"> <td align=\"center\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"https://www.epicypher.com/content/images/products/antibodies/13_0030_seq_tracks.jpg\" target=\"_new\"><img alt=\"13-0030 Sequencing Tracks\" height=\"100\" src=\"https://cdn11.bigcommerce.com/s-y9o92/content/images/products/antibodies/13_0030_seq_tracks.jpg\" width=\"127\" /></a></span></td> </tr> <tr> <td align=\"left\" valign=\"top\"> <p><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>H3K27me3 SNAP-ChIP-seq and CUT&RUN representative tracks: </strong>A gene browser shot generated using the Integrative Genomics Viewer (IGV, Broad Institute) shows a representative locus for EpiCypher H3K27me3 ChIP-seq replicate experiments (blue tracks, 5 μg antibody) and CUT&RUN (green track, 1:100 antibody dilution). For comparison ENCODE H3K27me3 ChIP-seq using a different antibody is shown (bottom orange track, GEO accession number GSM733658). Similar results in peak structure and location were observed throughout the genome for EpiCypher H3K27me3 antibody in ChIP-seq and CUT&RUN. <i>Methods: Native ChIP-seq was performed as described (Shah et al., Mol Cell 2018). CUT&RUN was performed using EpiCypher CUTANA pAG-MNase for ChIC/CUT&RUN (EpiCypher Catalog No. 15-1016) as described (EpiCypher.com/cutana-protocol). Library preparation was performed with 10 ng DNA using the NEBNext® UltraTM II DNA Library Prep Kit for Illumina®. ChIP libraries were sequenced on an Illumina NextSeq 550 (2x150bp paired end). The total number of reads was 37.8 and 34.9 million for the ChIP-seq replicates and 8.0 million for CUT&RUN.</i><br /> <strong>(Click image to enlarge) </strong></span></p> <p></p> </td> </tr>\n \n <tr valign=\"top\"> <td align=\"center\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"https://www.epicypher.com/content/images/products/antibodies/13_0030_tss.jpg\" target=\"_new\"><img alt=\"13-0030 Genome Wide Analysis\" height=\"100\" src=\"https://cdn11.bigcommerce.com/s-y9o92/content/images/products/antibodies/13_0030_tss.jpg\" width=\"127\" /></a></span></td> </tr> <tr> <td align=\"left\" valign=\"top\"> <p><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>ChIP-seq and CUT&RUN genome wide analysis: </strong>EpiCypher H3K27me3 antibody was tested in native ChIP-seq <strong> A </strong> and CUT&RUN <strong> B </strong> using the methods described above. Genome-wide analysis of H3K27me3 enrichment (signal intensity) flanking annotated transcription start sits (TSSs; +/- 3kb) is graphed as a cumulative histogram plot (top) and shown in a heatmap (bottom). Individual gene loci in each row of the heatmap are colored by signal intensity and sorted by strongest to lowest enrichment (top to bottom). EpiCypher H3K27me3 an2body displays a characteristic enrichment pattern downstream of the TSS, remaining elevated throughout the body of the gene.<br /> <strong>(Click image to enlarge) </strong></span></p> <p></p> </td> </tr> \n \n <tr valign=\"top\"> <td align=\"center\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"https://www.epicypher.com/content/images/products/antibodies/13_0030_ca_tracks.jpg\" target=\"_new\"><img alt=\"13-0030 Correlation Analysis\" height=\"100\" src=\"https://cdn11.bigcommerce.com/s-y9o92/content/images/products/antibodies/13_0030_ca_tracks.jpg\" width=\"127\" /></a></span></td> </tr> <tr> <td align=\"left\" valign=\"top\"> <p><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>ChIP-seq vs. CUT&RUN correlation analysis: </strong>Genome-wide correlation analysis was performed to compare EpiCypher H3K27me3 antibody enrichment in ChIP-seq and CUT&RUN. The number of reads per 10 kb binned region across the genome is plotted for CUT&RUN (x-axis) vs. ChIP-seq (y-axis) (EaSeq). ChIP-seq and CUT&RUN data generated using this antibody are highly correlated (Pearson correlation r = 0.864).<br /> <strong>(Click image to enlarge) </strong></span></p> <p></p> </td> </tr> \n \n <tr valign=\"top\"> <td align=\"center\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"https://www.epicypher.com/content/images/products/antibodies/13_0030_wb_tds.jpg\" target=\"_new\"><img alt=\"13-0030 Western Blot\" height=\"100\" src=\"https://cdn11.bigcommerce.com/s-y9o92/content/images/products/antibodies/13_0030_wb_tds.jpg\" width=\"127\" /></a></span></td> </tr> <tr> <td align=\"left\" valign=\"top\"> <p><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Western Blot Analysis: </strong>Recombinant histone H3.3 (Lane 1) and acid extracts of HeLa cells (Lane 2) were blotted onto PVDF and probed with 1 µg/mL EpiCypher H3K27me3 Antibody.<br /> <strong>(Click image to enlarge) </strong></span></p> <p></p> </td> </tr>\n \n <tr valign=\"top\"> <td align=\"center\" valign=\"top\"><span style=\"font-family:arial,helvetica,sans-serif;\"><a href=\"https://www.epicypher.com/content/images/products/antibodies/13_0030_IHC_tds.jpg\" target=\"_new\"><img alt=\"13-0030 Immunohistochemistry\" height=\"100\" src=\"https://cdn11.bigcommerce.com/s-y9o92/content/images/products/antibodies/13_0030_IHC_tds.jpg\" width=\"127\" /></a></span></td> </tr> <tr> <td align=\"left\" valign=\"top\"> <p><span style=\"font-family:arial,helvetica,sans-serif;\"><strong>Immunohistochemistry (IHC): </strong>IHC staining of HepG2 cells using EpiCypher\n H3K27me3\tAntibody.<br /> <strong>(Click image to enlarge) </strong></span></p> <p></p> </td> </tr></tbody> </table> </td> \n </tr> </tbody> </table> \n \n <style>\n .button-group {\n display: none;\n }\n </style>","tags":[],"detail_messages":"","availability":"","page_title":"Histone H3K27me3 Antibody, SNAP-ChIP® Certified, CUTANA™ CUT&RUN Compatible (13-0030)","mpn":"","upc":null,"options":[],"related_products":[{"id":411,"sku":null,"name":"SNAP-ChIP® K-MetStat Panel","url":"https://www.epicypher.com/products/nucleosomes/snap-chip-k-metstat-panel","availability":"","rating":null,"brand":{"name":null},"category":["Nucleosomes","Nucleosomes/SNAP-ChIP<sup>®</sup> Spike-ins"],"summary":"\n\t\n\t\t\n\t\t\t\n\t\t\t\n\t\t\t\t\n\t\t\t\t\t\n\t\t\t\t\t\t\n\t\t\t\t\t\t\n\n\t\t\t\t\t\t\n\t\t\t\t\t\tSNAP-ChIP K-MetStat Panel Description: A panel ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/411/282/SNAP_Chip__53961.1569012494.png?c=2","alt":"SNAP-ChIP® K-MetStat Panel"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/411/282/SNAP_Chip__53961.1569012494.png?c=2","alt":"SNAP-ChIP® K-MetStat Panel"}],"date_added":"18th Sep 2017","pre_order":false,"show_cart_action":true,"has_options":true,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":null,"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"price":{"without_tax":{"currency":"USD","formatted":"$365.00","value":365},"price_range":{"min":{"without_tax":{"currency":"USD","formatted":"$365.00","value":365},"tax_label":"Sales Tax"},"max":{"without_tax":{"currency":"USD","formatted":"$2,795.00","value":2795},"tax_label":"Sales Tax"}},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=411"}],"shipping_messages":[],"rating":0,"title":"Histone H3K27me3 Antibody, SNAP-ChIP® Certified, CUTANA™ CUT&RUN Compatible *DISCONTINUED*","gift_wrapping_available":false,"min_purchase_quantity":0} Pack Size: 100 μg
|
Current stock:
0
Related Products

$365.00