EpiDyne-FRET Nucleosome Remodeling Assay Substrate
{"url":"https://www.epicypher.com/products/nucleosomes/epidyne-fret-nucleosome-remodeling-assay-substrate","add_this":[{"service":"facebook","annotation":""},{"service":"email","annotation":""},{"service":"print","annotation":""},{"service":"twitter","annotation":""},{"service":"linkedin","annotation":""}],"gtin":"","id":"386","bulk_discount_rates":[],"can_purchase":true,"meta_description":"EpiDyne FRET-based Chromatin Remodeling Assay Substrate for inhibitor screening and HTS assays","category":["Nucleosomes","Nucleosomes/EpiDyne® Chromatin Remodeling Substrates"],"AddThisServiceButtonMeta":"","main_image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/386/276/2_EpiDyne__20008.1569012494.png?c=2","alt":"EpiDyne-FRET Nucleosome Remodeling Assay Substrate"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=386","shipping":{"calculated":true},"num_reviews":0,"weight":"0.00 LBS","custom_fields":[{"id":"1138","name":"Pack Size","value":"50 µg"},{"id":"1139","name":"Internal Comment","value":"Bulk in Bulk Nuc Box 2 in Psylocke"}],"sku":"16-4201","description":"<div class=\"service_accordion product-droppdown\">\n <div class=\"container\">\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel current\">\n <h3 class=\"sub-title1\">Description</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\"\n style=\"display: block\">\n <p>\n This product provides a high throughput fluorescent readout of\n nucleosome remodeling when paired with enzymatically active\n remodeling complexes, such as SMARCA4, SMARCA2, and ACF (EpiCypher\n <a\n href=\"/products/nucleosomes/epidyne-chromatin-remodeling-substrates/epidyne-modifying-enzymes/smarca4-chromatin-remodeling-enzyme-human-brg1\"\n >15-1014</a\n >,\n <a\n href=\"/products/nucleosomes/epidyne-chromatin-remodeling-substrates/epidyne-modifying-enzymes/smarca2-chromatin-remodeling-enzyme-human-brm\"\n >15-1015</a\n >,\n <a\n href=\"/products/nucleosomes/epidyne-chromatin-remodeling-substrates/epidyne-modifying-enzymes/acf-chromatin-remodeling-enzyme-complex\"\n >15-1013</a\n >). The EpiDyne<sup>®</sup>-FRET mononucleosome is assembled from\n recombinant human histones expressed in <em>E. coli</em> (two each\n of histones H2A-Cy5, H2B, H3.2 and H4; accession numbers:\n H2A-P04908; H2B-O60814; H3.2-Q71DI3; H4-P62805) wrapped with a 212\n base-pair DNA template bearing a 5’ Cy3, the Widom 601 nucleosome\n positioning element [1], and a DpnII restriction enzyme motif (GATC)\n that is exposed upon remodeling. H2A has a Thr to Cys substitution\n at residue 120, where Cy5 is conjugated. H3.2 has a Cys to Ala\n substitution at residue 110.\n </p>\n <p\n style=\"\n background-color: #4698cb;\n color: #fff;\n padding: 1.3rem;\n text-align: center;\n border-radius: 12px;\n \">\n <a\n target=\"_blank\"\n style=\"color: #fff\"\n href=\"/resources/protocols/epidyne-guide\"\n >Want to learn more? Check out our EpiDyne® Guide >>\n </a>\n </p>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel current\">\n <h3 class=\"sub-title1\">Validation Data</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\"\n style=\"display: block\">\n <section class=\"image-picker\">\n <div class=\"image-picker__left\">\n <div\n class=\"image-picker__main-content_active image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg\n class=\"image-picker__svg-left\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\">\n <path\n d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a\n href=\"content/images/products/nucleosomes/16-4201-dna-gel-data.jpeg\"\n target=\"_new\"\n class=\"image-picker__main-image-link\"\n ><img loading=\"lazy\"\n alt=\"16-4201-dna-gel-data\"\n src=\"/content/images/products/nucleosomes/16-4201-dna-gel-data.jpeg\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\"\n >(Click to enlarge)</span\n ></a\n >\n <button class=\"image-picker__right-arrow\">\n <svg\n class=\"image-picker__svg-right\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\">\n <path\n d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\"\n ><strong>Figure 1: DNA Gel Data </strong>\n <br />\n EpiDyne-FRET substrate resolved via native PAGE gel and\n stained with ethidium bromide to visualize DNA.\n <strong>Lane 1</strong>: Free DNA (100 ng).\n <strong>Lane 2</strong>: Intact EpiDyne-FRET nucleosomes\n (400 ng).\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg\n class=\"image-picker__svg-left\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\">\n <path\n d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a\n href=\"/content/images/products/nucleosomes/16-4201-protein-gel-data.jpeg\"\n target=\"_new\"\n class=\"image-picker__main-image-link\"\n ><img loading=\"lazy\"\n alt=\"16-4201-protein-gel-data\"\n src=\"/content/images/products/nucleosomes/16-4201-protein-gel-data.jpeg\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\"\n >(Click to enlarge)</span\n ></a\n >\n <button class=\"image-picker__right-arrow\">\n <svg\n class=\"image-picker__svg-right\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\">\n <path\n d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\"\n ><strong>Figure 2: Protein Gel Data </strong><br />\n Coomassie stained PAGE gel of proteins in EpiDyne-FRET\n substrate (1 µg) demonstrates the purity of histones in the\n preparation. Sizes of molecular weight markers and positions\n of the core histones (H2A-Cy5, H2B, H3.2, and H4) are\n indicated.\n </span>\n </p>\n </div>\n <div class=\"image-picker__main-content\">\n <div class=\"image-picker__header-content\">\n <button class=\"image-picker__left-arrow\">\n <svg\n class=\"image-picker__svg-left\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\">\n <path\n d=\"M16.67 0l2.83 2.829-9.339 9.175 9.339 9.167-2.83 2.829-12.17-11.996z\" />\n </svg>\n </button>\n <a\n href=\"/content/images/products/nucleosomes/16-4201-nucleosome-remodeling-data.jpeg\"\n target=\"_new\"\n class=\"image-picker__main-image-link\">\n <img loading=\"lazy\"\n alt=\"16-4201-nucleosome-remodeling-data\"\n src=\"/content/images/products/nucleosomes/16-4201-nucleosome-remodeling-data.jpeg\"\n class=\"image-picker__main-image\" />\n <span class=\"image-picker__main-image-caption\"\n >(Click to enlarge)</span\n >\n </a>\n <button class=\"image-picker__right-arrow\">\n <svg\n class=\"image-picker__svg-right\"\n width=\"24\"\n height=\"24\"\n viewBox=\"0 0 24 24\">\n <path\n d=\"M7.33 24l-2.83-2.829 9.339-9.175-9.339-9.167 2.83-2.829 12.17 11.996z\" />\n </svg>\n </button>\n </div>\n <p>\n <span class=\"image-picker__span-content\">\n <strong>Figure 3: Nucleosome Remodeling Data </strong><br />\n ACF1/ATP-dependent nucleosome remodeling reaction.\n EpiDyne-FRET nucleosomes (15 nM) were incubated with ACF1\n chromatin remodeler (EpiCypher\n <a\n href=\"/products/nucleosomes/epidyne-chromatin-remodeling-substrates/epidyne-modifying-enzymes/acf-chromatin-remodeling-enzyme-complex\"\n >15-1013</a\n >; indicated concentrations) in the presence or absence of 1\n mM ATP. Upon the addition of ATP, reactions were immediately\n read in an Envision Multilabel plate reader (PerkinElmer).\n Data are presented as the mean of the Cy3/Cy5 ratio (N=2).\n </span>\n </p>\n </div>\n </div>\n <aside class=\"image-picker__right\">\n <div class=\"image-picker__gallery\">\n <img loading=\"lazy\"\n alt=\"16-4201-dna-gel-data\"\n src=\"/content/images/products/nucleosomes/16-4201-dna-gel-data.jpeg\"\n width=\"200\"\n class=\"image-picker__side-image image-picker__side-image_active\"\n role=\"button\" />\n <img loading=\"lazy\"\n alt=\"16-4201-protein-gel-data\"\n src=\"/content/images/products/nucleosomes/16-4201-protein-gel-data.jpeg\"\n class=\"image-picker__side-image\"\n role=\"button\" />\n <img loading=\"lazy\"\n alt=\"16-4201-nucleosome-remodeling-data\"\n src=\"/content/images/products/nucleosomes/16-4201-nucleosome-remodeling-data.jpeg\"\n class=\"image-picker__side-image\"\n role=\"button\" />\n </div>\n </aside>\n </section>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Technical Information</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\">\n <div class=\"product-tech-info\">\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Formulation</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n <p>\n Purified recombinant mononucleosomes, (21.8 µg protein weight,\n 50 µg DNA+protein) in 10 mM Tris-HCl pH 7.5, 25 mM NaCl, 1 mM\n EDTA, 2 mM DTT, 20% glycerol. MW = 240,650 Da.\n </p>\n </div>\n </div>\n <div class=\"product-tech-info__line-item\">\n <div class=\"product-tech-info__line-item-left\">\n <b>Storage and Stability</b>\n </div>\n <div class=\"product-tech-info__line-item-right\">\n Stable for six months at -80°C from date of receipt. For\n best results, aliquot and avoid multiple freeze/thaws.\n </div>\n </div>\n </div>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Application Notes</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\">\n <p>\n This product is a template for nucleosome remodeling assays using\n Cy3/Cy5 FRET or using the restriction enzyme DpnII to determine\n accessibility of GATC, which is masked in its native configuration\n (prior to remodeling).\n </p>\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">References</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\">\n <strong>Background References:</strong>\n <br />\n [1] Lowary & Widom <em>J Mol. Biol.</em> (1998) PMID:\n <a\n href=\"https://pubmed.ncbi.nlm.nih.gov/9514715/\"\n target=\"_blank\"\n title=\"New DNA sequence rules for high affinity binding to histone octamer and sequence-directed nucleosome positioning\"\n >9514715</a\n ><br />\n </div>\n </div>\n </div>\n <div id=\"prodAccordion\">\n <div id=\"ProductDescription\" class=\"Block Panel\">\n <h3 class=\"sub-title1\">Documents & Resources</h3>\n <div\n class=\"ProductDescriptionContainer product-droppdown__section-description\">\n <div class=\"product-documents\">\n <a\n href=\"/content/documents/tds/16-4201.pdf\"\n target=\"_new\"\n class=\"product-documents__link\">\n <svg\n version=\"1.1\"\n id=\"Layer_1\"\n xmlns=\"http://www.w3.org/2000/svg\"\n xmlns:xlink=\"http://www.w3.org/1999/xlink\"\n x=\"0px\"\n y=\"0px\"\n viewBox=\"0 0 228 240\"\n style=\"enable-background: new 0 0 228 240\"\n xml:space=\"preserve\"\n class=\"product-documents__icon\">\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M191.92,68.77l-47.69-47.69c-1.33-1.33-3.12-2.08-5.01-2.08H45.09C41.17,19,38,22.17,38,26.09v184.36\n c0,3.92,3.17,7.09,7.09,7.09h141.82c3.92,0,7.09-3.17,7.09-7.09V73.8C194,71.92,193.25,70.1,191.92,68.77z M177.65,77.06h-41.7\n v-41.7L177.65,77.06z M178.05,201.59H53.95V34.95h66.92v47.86c0,5.14,4.17,9.31,9.31,9.31h47.86V201.59z\" />\n </g>\n <rect\n x=\"20\"\n y=\"112\"\n class=\"product-documents__svg-background\"\n width=\"146\"\n height=\"76\" />\n <g>\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M23.83,125.68h22.36c5.29,0,9.41,1.33,12.35,4c2.94,2.67,4.42,6.39,4.42,11.18c0,4.78-1.47,8.51-4.42,11.18\n c-2.94,2.67-7.06,4-12.35,4H34.59v18.29H23.83V125.68z M44.81,147.9c5.38,0,8.07-2.32,8.07-6.97c0-2.39-0.67-4.16-2-5.31\n c-1.33-1.15-3.36-1.73-6.07-1.73H34.59v14.01H44.81z\" />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M69.92,125.68h18.91c5.29,0,9.84,0.97,13.66,2.9c3.82,1.93,6.74,4.72,8.76,8.35\n c2.02,3.63,3.04,7.98,3.04,13.04c0,5.06-1,9.42-3,13.08c-2,3.66-4.91,6.45-8.73,8.38c-3.82,1.93-8.4,2.9-13.73,2.9H69.92V125.68z\n M88.07,165.63c10.35,0,15.52-5.22,15.52-15.66c0-10.4-5.17-15.59-15.52-15.59h-7.38v31.26H88.07z\" />\n <path\n class=\"product-documents__svg-pdf\"\n d=\"M122.57,125.68h32.84v8.49h-22.22v11.18h20.84v8.49h-20.84v20.49h-10.63V125.68z\" />\n </g>\n </svg>\n <span class=\"product-documents__info\">Technical Datasheet</span>\n </a>\n </div>\n </div>\n </div>\n </div>\n </div>\n</div>\n","tags":[],"warranty":"","price":{"without_tax":{"formatted":"$595.00","value":595,"currency":"USD"},"tax_label":"Sales Tax"},"detail_messages":"","availability":"","page_title":"EpiDyne FRET-based Chromatin Remodeling Assay Substrate","cart_url":"https://www.epicypher.com/cart.php","max_purchase_quantity":0,"mpn":"","upc":null,"options":[],"related_products":[{"id":650,"sku":"15-1015","name":"SMARCA2 Chromatin Remodeling Enzyme (Human BRM)","url":"https://www.epicypher.com/products/nucleosomes/epidyne-chromatin-remodeling-substrates/epidyne-modifying-enzymes/smarca2-chromatin-remodeling-enzyme-human-brm","availability":"","rating":null,"brand":{"name":null},"category":["Nucleosomes/EpiDyne® Chromatin Remodeling Substrates/EpiDyne<sup>®</sup> Modifying Enzymes ","Recombinant Proteins/EpiDyne<sup>®</sup> Chromatin Remodeling Enzymes","Recombinant Proteins/Modifying Enzymes"],"summary":"\n \n \n Species: Human\n \n \n Source: SF9 cells\n \n \n \n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/650/953/smarca2-chromatin-remodeling-enzyme-human-brm__84445.1645734545.jpg?c=2","alt":"SMARCA2 Chromatin Remodeling Enzyme Human BRM"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/650/953/smarca2-chromatin-remodeling-enzyme-human-brm__84445.1645734545.jpg?c=2","alt":"SMARCA2 Chromatin Remodeling Enzyme Human BRM"}],"date_added":"15th Feb 2019","pre_order":false,"show_cart_action":true,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":1154,"name":"Pack Size","value":"100 Reactions"},{"id":1155,"name":"Internal Comment","value":"extra in top right of Psylocke"},{"id":1156,"name":"Internal Comment","value":"bulk in LG"}],"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"add_to_cart_url":"https://www.epicypher.com/cart.php?action=add&product_id=650","price":{"without_tax":{"currency":"USD","formatted":"$575.00","value":575},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=650"},{"id":651,"sku":"15-1014","name":"SMARCA4 Chromatin Remodeling Enzyme (Human BRG1)","url":"https://www.epicypher.com/products/nucleosomes/epidyne-chromatin-remodeling-substrates/epidyne-modifying-enzymes/smarca4-chromatin-remodeling-enzyme-human-brg1","availability":"","rating":null,"brand":{"name":null},"category":["Nucleosomes/EpiDyne® Chromatin Remodeling Substrates/EpiDyne<sup>®</sup> Modifying Enzymes ","Recombinant Proteins/EpiDyne<sup>®</sup> Chromatin Remodeling Enzymes","Recombinant Proteins/Modifying Enzymes"],"summary":"\n \n \n Species: Human\n \n \n Source: SF9 cells\n \n \n \n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/651/616/Untitled__68998.1556128740.png?c=2","alt":"SMARCA4 Chromatin Remodeling Enzyme (Human BRG1)"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/651/616/Untitled__68998.1556128740.png?c=2","alt":"SMARCA4 Chromatin Remodeling Enzyme (Human BRG1)"}],"date_added":"15th Feb 2019","pre_order":false,"show_cart_action":true,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":1140,"name":"Pack size","value":"100 reactions"},{"id":1141,"name":"Internal Comment","value":"21-1014 is sold with this item."},{"id":1142,"name":"Internal Comment","value":"51 units of R&D Lot 7 are in HQ"},{"id":1143,"name":"Internal Comment","value":"Bulk in Protein Box in LG, Lot 7"}],"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"add_to_cart_url":"https://www.epicypher.com/cart.php?action=add&product_id=651","price":{"without_tax":{"currency":"USD","formatted":"$575.00","value":575},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=651"},{"id":551,"sku":"15-1013","name":"ACF Chromatin Remodeling Enzyme Complex","url":"https://www.epicypher.com/products/nucleosomes/epidyne-chromatin-remodeling-substrates/epidyne-modifying-enzymes/acf-chromatin-remodeling-enzyme-complex","availability":"","rating":null,"brand":{"name":null},"category":["Nucleosomes/EpiDyne® Chromatin Remodeling Substrates/EpiDyne<sup>®</sup> Modifying Enzymes ","Recombinant Proteins/EpiDyne<sup>®</sup> Chromatin Remodeling Enzymes","Recombinant Proteins/Modifying Enzymes"],"summary":"\n \n \n \n \n \n \n \n \n \n \n \n \n \n \n \n \n \n ","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/551/597/Untitled__22497.1556128753.png?c=2","alt":"ACF Chromatin Remodeling Enzyme Complex"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/551/597/Untitled__22497.1556128753.png?c=2","alt":"ACF Chromatin Remodeling Enzyme Complex"}],"date_added":"24th Jul 2018","pre_order":false,"show_cart_action":true,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":585,"name":"Pack Size","value":"100 reactions"}],"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"add_to_cart_url":"https://www.epicypher.com/cart.php?action=add&product_id=551","price":{"without_tax":{"currency":"USD","formatted":"$505.00","value":505},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=551"},{"id":347,"sku":"16-4101","name":"EpiDyne® Nucleosome Remodeling Assay Substrate ST601-GATC1","url":"https://www.epicypher.com/products/nucleosomes/epidyne-nucleosome-remodeling-assay-substrate-st601-gatc1","availability":"","rating":null,"brand":{"name":null},"category":["Nucleosomes","Nucleosomes/EpiDyne® Chromatin Remodeling Substrates"],"summary":"\n \n \n \n Description\n \n \n Mononucleosomes assembled","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/347/803/2021.02.25_EPI_Nucleosome_thumbnail_only_6N66_white_background_RGB_AL_MP__47518.1620836275.png?c=2","alt":"EpiDyne® Nucleosome Remodeling Assay Substrate ST601-GATC1"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/347/803/2021.02.25_EPI_Nucleosome_thumbnail_only_6N66_white_background_RGB_AL_MP__47518.1620836275.png?c=2","alt":"EpiDyne® Nucleosome Remodeling Assay Substrate ST601-GATC1"},{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/347/802/2021.02.25_EPI_Nucleosome_thumbnail_only_6N66_no_background_RGB_AL_MP__86463.1620836275.png?c=2","alt":"EpiDyne® Nucleosome Remodeling Assay Substrate ST601-GATC1"}],"date_added":"1st Mar 2017","pre_order":false,"show_cart_action":true,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":1053,"name":"Pack Size","value":"50 μg"},{"id":1054,"name":"Internal Comment","value":"no more bulk 3/31/22"}],"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"add_to_cart_url":"https://www.epicypher.com/cart.php?action=add&product_id=347","price":{"without_tax":{"currency":"USD","formatted":"$565.00","value":565},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=347"},{"id":556,"sku":"18-4201","name":"EpiDyne-FRET Remodeling Assay Substrate DNA","url":"https://www.epicypher.com/products/nucleosomes/epidyne-fret-remodeling-assay-substrate-dna","availability":"","rating":null,"brand":{"name":null},"category":["Nucleosomes","Nucleosomes/EpiDyne® Chromatin Remodeling Substrates"],"summary":"","image":{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/556/401/2_EpiDyne__74802.1566413754.png?c=2","alt":"EpiDyne-FRET Remodeling Assay Substrate DNA"},"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/556/401/2_EpiDyne__74802.1566413754.png?c=2","alt":"EpiDyne-FRET Remodeling Assay Substrate DNA"}],"date_added":"1st Aug 2018","pre_order":false,"show_cart_action":true,"has_options":false,"stock_level":null,"low_stock_level":null,"qty_in_cart":0,"custom_fields":[{"id":600,"name":"Pack Size","value":"50 µg"}],"num_reviews":null,"weight":{"formatted":"0.01 LBS","value":0.01},"demo":false,"add_to_cart_url":"https://www.epicypher.com/cart.php?action=add&product_id=556","price":{"without_tax":{"currency":"USD","formatted":"$165.00","value":165},"tax_label":"Sales Tax"},"add_to_wishlist_url":"/wishlist.php?action=add&product_id=556"}],"shipping_messages":[],"rating":0,"meta_keywords":"ATPase, EpiDyne-FRET Nucleosome Remodeling Assay Substrate, EpiDyne-FRET, chromatin remodeling, chromatin remodeler, swi.snf, swi, snf, chrac, rsc, SMARCA, BRM1, BRG1, 16-4201, nucleosome remodeling, nucleosome sliding, sliding, restriction enzyme assay, DAM methyltransferase assay, , chromatin remodeling, BRM, CHD, chromatin remodeling factors, SMARCA, , SMARCA4, SNF2L4, nucleosome positioning, SWR1, ISWI, BAF190 assay, BAF190 complex, BAF190 remodeler, BAF190A, BAF190A assay, BAF190A atpase, BAF190A complex, BAF190A remodeler, BAF190B, BAF190B assay, BAF190B atpase, BAF190B complex, BAF190B remodeler, BRG1 assay, BRG1 atpase, BRG1 complex, BRG1 remodeler, BRM assay, BRM atpase, BRM complex, BRM remodeler, CHD assay, CHD atpase, CHD complex, CHD remodeler, INO80, INO80 assay, INO80 atpase, INO80 complex, INO80 remodeler, ISWI assay, ISWI atpase, ISWI complex, ISWI remodeler, Mi-2, Mi-2 assay, Mi-2 atpase, Mi-2 complex, Mi-2 remodeler, NuRD, NuRD assay, NuRD atpase, NuRD complex, NuRD remodeler, SMARCA assay, SMARCA atpase, SMARCA complex, SMARCA remodeler, SMARCA4 assay, SMARCA4 atpase, SMARCA4 complex, SMARCA4 remodeler, SNF2A, SNF2A assay, SNF2A atpase, SNF2A complex, SNF2A remodeler, SNF2B, SNF2B assay, SNF2B atpase, SNF2B complex, SNF2B remodeler, SNF2L2, SNF2L2 assay, SNF2L2 atpase, SNF2L2 complex, SNF2L2 remodeler, SNF2L4 assay, SNF2L4 atpase, SNF2L4 complex, SNF2L4 remodeler, SWI/SNF, SWI/SNF assay, SWI/SNF atpase, SWI/SNF complex, SWI/SNF remodeler, SWR1 assay, SWR1 atpase, SWR1 complex, SWR1 remodeler, chromatin remodeler, nucleosome remodeler, nucleosome remodeling assay, chromatin remodeling assay, nucleosome remodeling, chromatin remodeling complex, nucleosome remodeling complex, BRG1, nucleosome sliding, nucleosome sliding assay, nucleosome atpase, nucleosome mobilization, FRET, EpiDyne Nucleosome Remodeling Assay Substrate, EpiDyne, chromatin remodeling","show_quantity_input":1,"title":"EpiDyne-FRET Nucleosome Remodeling Assay Substrate","gift_wrapping_available":false,"min_purchase_quantity":0,"customizations":[],"images":[{"data":"https://cdn11.bigcommerce.com/s-y9o92/images/stencil/{:size}/products/386/276/2_EpiDyne__20008.1569012494.png?c=2","alt":"EpiDyne-FRET Nucleosome Remodeling Assay Substrate"}]} Pack Size: 50 µg
Description
This product provides a high throughput fluorescent readout of nucleosome remodeling when paired with enzymatically active remodeling complexes, such as SMARCA4, SMARCA2, and ACF (EpiCypher 15-1014, 15-1015, 15-1013). The EpiDyne®-FRET mononucleosome is assembled from recombinant human histones expressed in E. coli (two each of histones H2A-Cy5, H2B, H3.2 and H4; accession numbers: H2A-P04908; H2B-O60814; H3.2-Q71DI3; H4-P62805) wrapped with a 212 base-pair DNA template bearing a 5’ Cy3, the Widom 601 nucleosome positioning element [1], and a DpnII restriction enzyme motif (GATC) that is exposed upon remodeling. H2A has a Thr to Cys substitution at residue 120, where Cy5 is conjugated. H3.2 has a Cys to Ala substitution at residue 110.
Validation Data
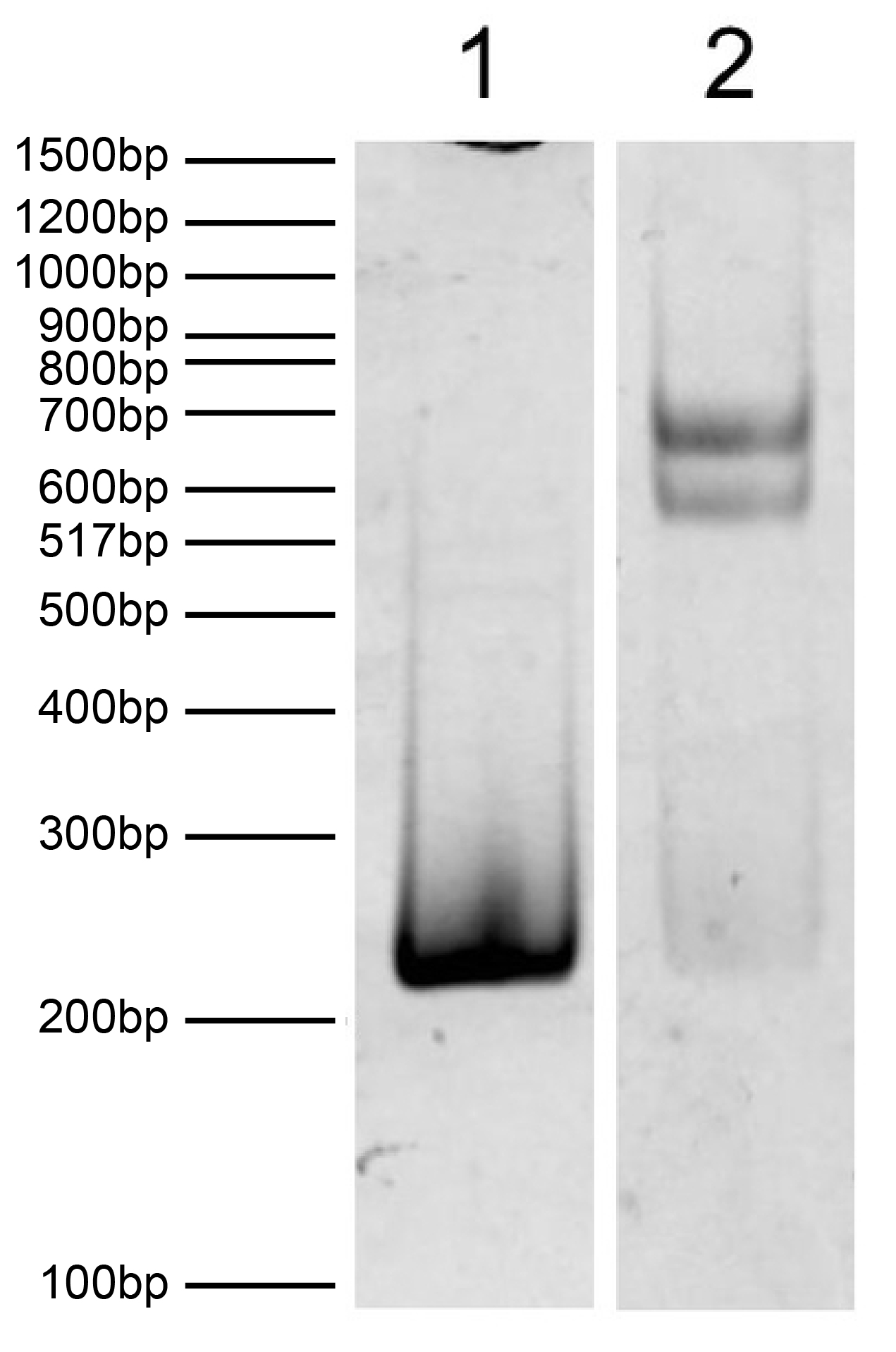
Figure 1: DNA Gel Data
EpiDyne-FRET substrate resolved via native PAGE gel and
stained with ethidium bromide to visualize DNA.
Lane 1: Free DNA (100 ng).
Lane 2: Intact EpiDyne-FRET nucleosomes
(400 ng).
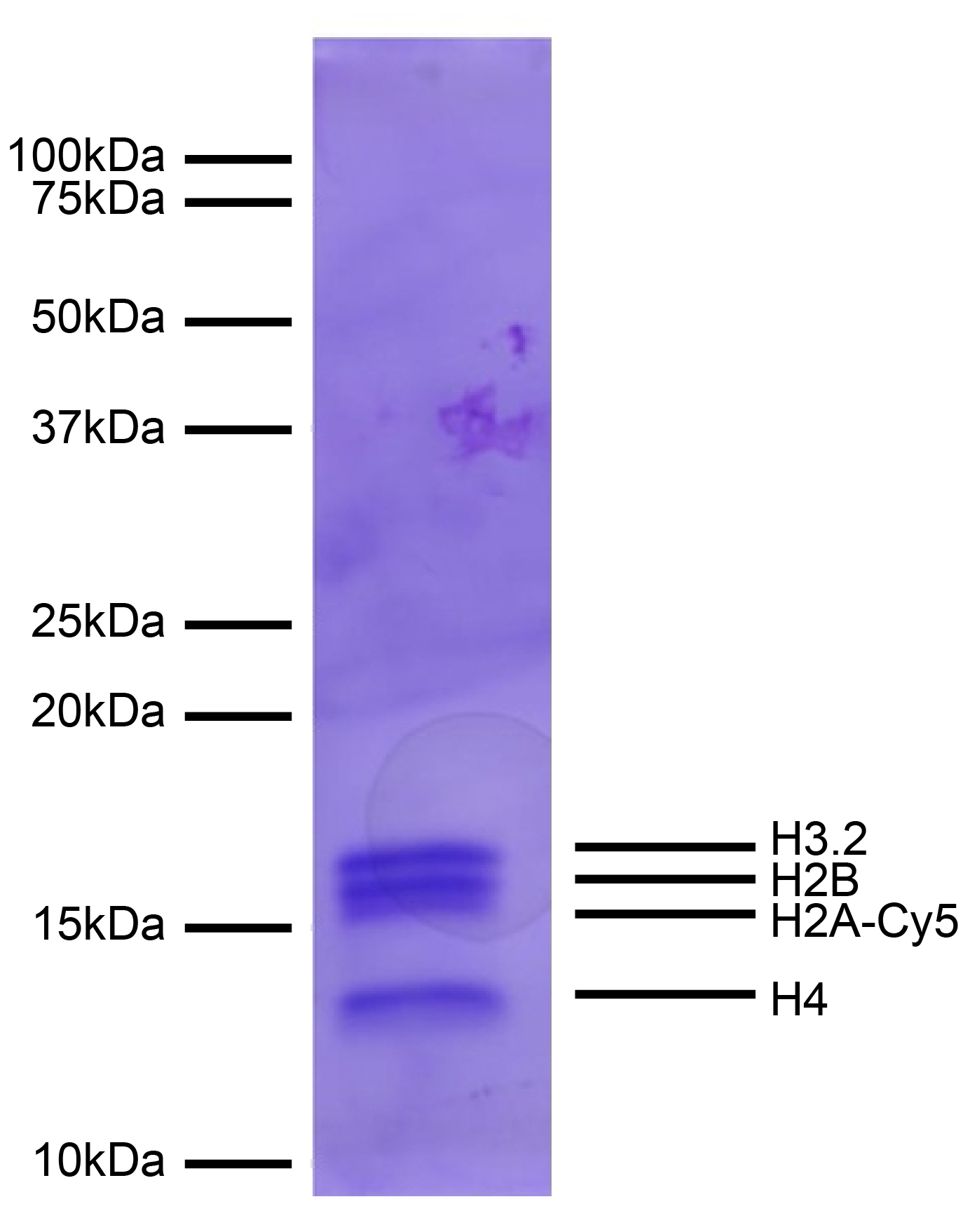
Figure 2: Protein Gel Data
Coomassie stained PAGE gel of proteins in EpiDyne-FRET
substrate (1 µg) demonstrates the purity of histones in the
preparation. Sizes of molecular weight markers and positions
of the core histones (H2A-Cy5, H2B, H3.2, and H4) are
indicated.
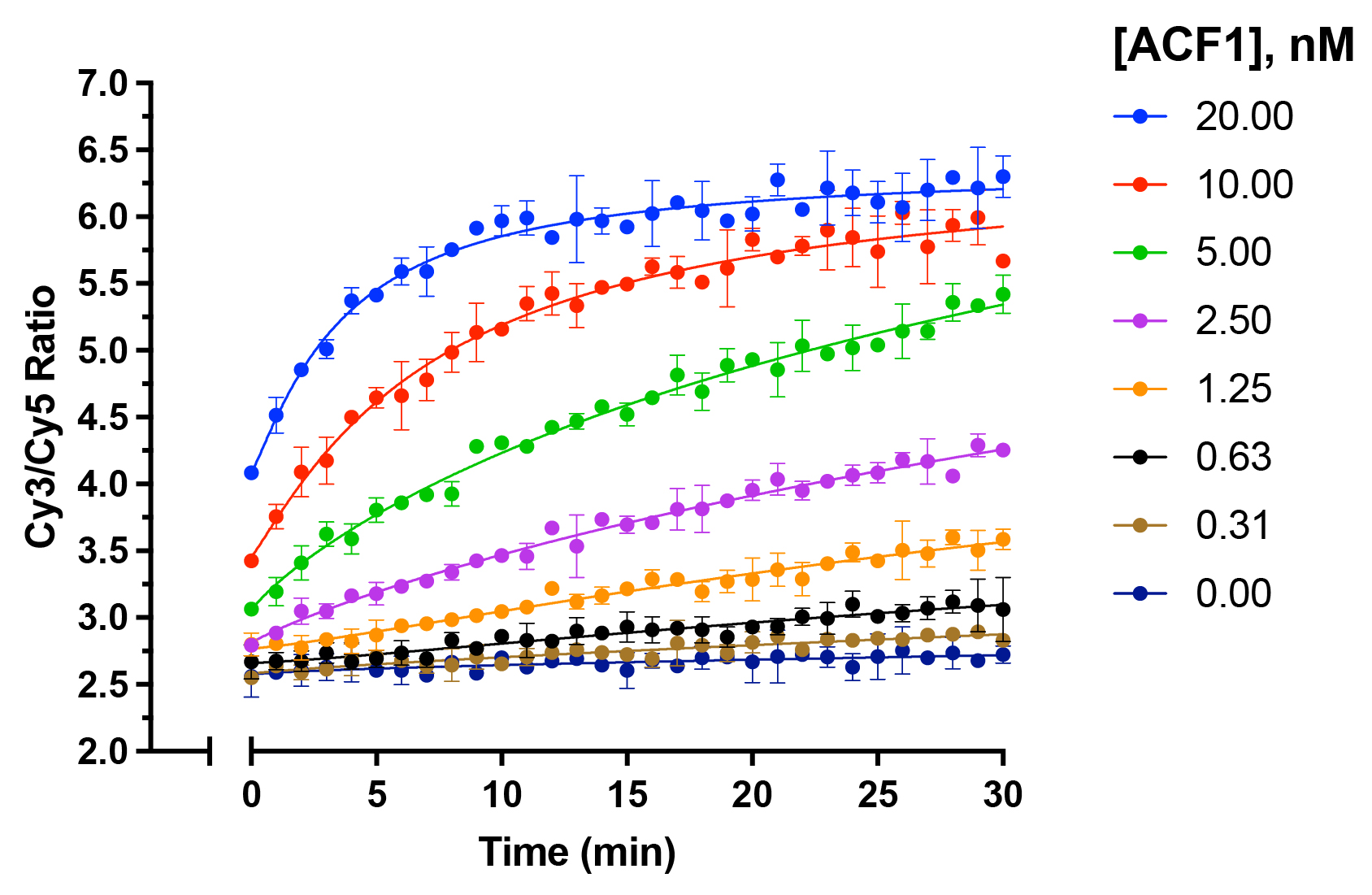
Figure 3: Nucleosome Remodeling Data
ACF1/ATP-dependent nucleosome remodeling reaction.
EpiDyne-FRET nucleosomes (15 nM) were incubated with ACF1
chromatin remodeler (EpiCypher
15-1013; indicated concentrations) in the presence or absence of 1
mM ATP. Upon the addition of ATP, reactions were immediately
read in an Envision Multilabel plate reader (PerkinElmer).
Data are presented as the mean of the Cy3/Cy5 ratio (N=2).
Technical Information
Purified recombinant mononucleosomes, (21.8 µg protein weight, 50 µg DNA+protein) in 10 mM Tris-HCl pH 7.5, 25 mM NaCl, 1 mM EDTA, 2 mM DTT, 20% glycerol. MW = 240,650 Da.
Application Notes
This product is a template for nucleosome remodeling assays using Cy3/Cy5 FRET or using the restriction enzyme DpnII to determine accessibility of GATC, which is masked in its native configuration (prior to remodeling).