Peptide or Nucleosome? dCypher™ Reveals the Optimal Substrate for Histone PTM Analyses
Histone ‘reader’ proteins contain evolutionarily conserved domains that bind to specific histone post-translational modifications (PTMs)1. These proteins have been shown to regulate diverse cellular processes via recruitment of epigenetic complexes or chromatin modifying / remodeling enzymes. Histone readers are an untapped wealth of information that underlie the foundation of chromatin regulation and could be used to advance therapeutics, diagnostics, and even personalized medicine. However, very little is known about how this diverse class of interactors functions in the context of chromatin.
Histone peptides:
an incomplete chromatin model?
For years, histone peptide-based assays have been the gold standard for developing antibodies against PTMs, as well as defining the targets of chromatin modifying enzymes and histone readers. Scientists have depended on this data to build essential theories of chromatin function, as it relates to gene expression, DNA damage repair, and the development of human disease.
It has largely been assumed that reader interactions with histone PTMs occur independently of the larger nucleosomal structure, supporting the use of peptides or peptide arrays in chromatin studies. However, recent NMR spectroscopy data reveal that histone PTMs display substantial interactions with nucleosomal DNA, and that these contacts influence protein binding activity2-5. Furthermore, EpiCypher’s own SNAP-ChIP® (Sample Normalization and Antibody Profiling for ChIP) platform has shown that histone peptides fail to accurately predict antibody binding specificity in ChIP experiments6. To learn more, see our recent Molecular Cell publication6.
All of this points to a misguided dependence on peptide-based data (such as histone peptide arrays), but direct comparison of peptides and nucleosomal substrates in chromatin reader assays has been difficult to achieve.
dCypher as an innovative platform to test
histone peptides vs. nucleosomes
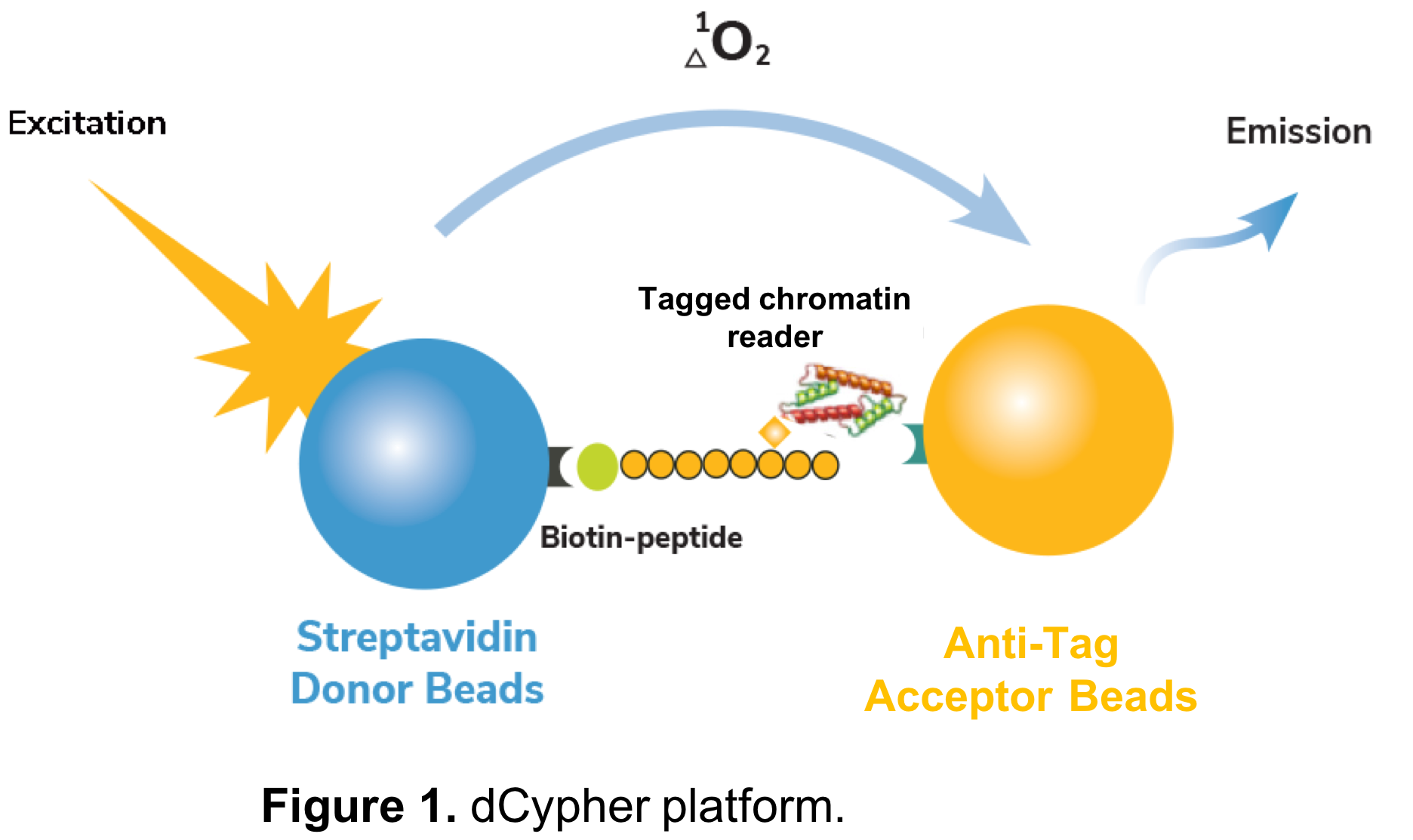 This is where EpiCypher’s recently released dCypher™ services come into play. Our dCypher approach is based on a no-wash, bead-based proximity assay (AlphaScreen®), and was originally designed to interrogate chromatin reader domains against a library of singly- or combinatorially-modified histone peptides (Figure 1). dCypher pairs biotinylated modified histone peptides to streptavidin-coated Donor beads, and tagged chromatin reader domains are paired with anti-tag Acceptor beads. Interaction of reader domains with dNucs brings the Donor and Acceptor beads into close proximity, enabling light-induced excitation of the Acceptor bead, which is released in the form of fluorescence (Figure 1). The amount of light emitted is directly proportional to the quantity of chromatin reader-dNuc binding. As many chromatin reader domain targets are ill-defined, this platform was developed to reveal novel interactions and guide drug target / inhibitor screens.
This is where EpiCypher’s recently released dCypher™ services come into play. Our dCypher approach is based on a no-wash, bead-based proximity assay (AlphaScreen®), and was originally designed to interrogate chromatin reader domains against a library of singly- or combinatorially-modified histone peptides (Figure 1). dCypher pairs biotinylated modified histone peptides to streptavidin-coated Donor beads, and tagged chromatin reader domains are paired with anti-tag Acceptor beads. Interaction of reader domains with dNucs brings the Donor and Acceptor beads into close proximity, enabling light-induced excitation of the Acceptor bead, which is released in the form of fluorescence (Figure 1). The amount of light emitted is directly proportional to the quantity of chromatin reader-dNuc binding. As many chromatin reader domain targets are ill-defined, this platform was developed to reveal novel interactions and guide drug target / inhibitor screens.
Subscribe for the Latest in Chromatin Technology!
Although we have used dCypher to identify novel reader interactions using our expansive modified histone peptide collection, EpiCypher is constantly thinking about how to apply and innovate our platforms to address important problems in epigenetics. This is precisely what we have done with our dCypher platform. By incorporating biotinylated recombinant designer nucleosomes carrying physiological histone PTMs (aka dNucs), we have created a system that finally allows rapid head-to-head comparison of histone peptide and nucleosome binding to chromatin readers. For example, we have applied this innovative system to the study of the HP1β chromodomain (CHD), which is reported to have selectivity for H3K9me1, me2, and me3 histone peptides7. We confirmed this selectivity using our standard histone-peptide based dCypher assay (Figure 2A), but were curious as to whether HP1β CHD specificity would differ in a nucleosomal context.
Nucleosomal context alters
HP1β reader specificity

We were both surprised and impressed with the results (shown in Figure 2B). When presented with H3K9me1, H3K9me2, and H3K9me3 marked dNucs, HP1β CHD showed exceptional specificity for H3K9me3 (Figure 2B). This is in direct contrast to our own histone peptide data (Figure 2A) as well as previously published reports7. The nucleosomal context is clearly impacts substrate selectivity of the HP1β chromatin reader domain; however, the mechanism by which this occurs remains unclear8. Using this experimental design, EpiCypher is currently testing additional histone reader domains to determine if this is a common feature of reader motifs, or if it is specific to certain protein families or subtypes.
But, what does this mean for the chromatin field? What is the impact of this discovery in other areas of epigenetics research?
The most important conclusion to draw from this work is that we are entering an exciting period of technological advancement, with new tools that will both confirm and correct our preconceived notions of chromatin dynamics. We have depended on histone peptides for so long because they were, essentially, our only option. Now, by using EpiCypher’s cutting edge dNuc-based technologies (such as dCypher, described here) we have the potential to rapidly and dramatically redefine our fundamental understanding of chromatin biology regulation.
References
1. Musselman CA, et al. Perceiving the epigenetic landscape through histone readers. Nat Struct Mol Biol, 2012. 19(12): p. 1218-27. (PubMed PMID: 23211769) (PMC3645987)
2. Morrison EA, et al. The conformation of the histone H3 tail inhibits association of the BPTF PHD finger with the nucleosome. Elife, 2018. 7: p. (PubMed PMID: 29648537) (PMC5953545)
3. Weaver TM, et al. Reading More than Histones: The Prevalence of Nucleic Acid Binding among Reader Domains. Molecules, 2018. 23(10): p. (PubMed PMID: 30322003) (PMC6222470)
4. Gatchalian J, et al. Accessibility of the histone H3 tail in the nucleosome for binding of paired readers. Nat Commun, 2017. 8(1): p. 1489. (PubMed PMID: 29138400) (PMC5686127)
5. Morrison EA, et al. DNA binding drives the association of BRG1/hBRM bromodomains with nucleosomes. Nat Commun, 2017. 8: p. 16080. (PubMed PMID: 28706277) (PMC5519978)
6. Shah RN, et al. Examining the Roles of H3K4 Methylation States with Systematically Characterized Antibodies. Mol Cell, 2018. 72(1): p. 162-77 e7. (PubMed PMID: 30244833) (PMC6173622)
7. Kim J, et al. Tudor, MBT and chromodomains gauge the degree of lysine methylation. EMBO Rep, 2006. 7(4): p. 397-403. (PubMed PMID: 16415788) (PMC1456902)
8. Lalonde ME, et al. Histone target selection within chromatin: an exemplary case of teamwork. Genes Dev, 2014. 28(10): p. 1029-41. (PubMed PMID: 24831698) (PMC4035532)
